The marburg virus 3' noncoding region structurally and functionally differs from that of ebola virus
- PMID: 19225002
- PMCID: PMC2668471
- DOI: 10.1128/JVI.02429-08
The marburg virus 3' noncoding region structurally and functionally differs from that of ebola virus
Abstract
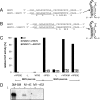
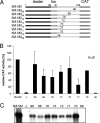
We have previously shown that the first transcription start signal (TSS) of Zaire Ebola virus (ZEBOV) is involved in formation of an RNA secondary structure regulating VP30-dependent transcription activation. Interestingly, transcription of Marburg virus (MARV) minigenomes occurs independently of VP30. In this study, we analyzed the structure of the MARV 3' noncoding region and its influence on VP30 necessity. Secondary structure formation of the TSS of the first gene was experimentally determined and showed substantial differences from the structure formed by the ZEBOV TSS. Chimeric MARV minigenomes mimicking the ZEBOV-specific RNA secondary structure were neither transcribed nor replicated. Mapping of the MARV genomic replication promoter revealed that the region homologous to the sequence involved in formation of the regulatory ZEBOV RNA structure is part of the MARV promoter. The MARV promoter is contained within the first 70 nucleotides of the genome and consists of two elements separated by a spacer region, comprising the TSS of the first gene. Mutations within the spacer abolished transcription activity and led to increased replication, indicating competitive transcription and replication initiation. The second promoter element is located within the nontranslated region of the first gene and consists of a stretch of three UN(5) hexamers. Recombinant full-length MARV clones, in which the three conserved U residues were substituted, could not be rescued, underlining the importance of the UN(5) hexamers for replication activity. Our data suggest that differences in the structure of the genomic replication promoters might account for the different transcription strategies of Marburg and Ebola viruses.
Figures
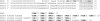



Similar articles
-
Phosphorylated VP30 of Marburg Virus Is a Repressor of Transcription.J Virol. 2018 Oct 12;92(21):e00426-18. doi: 10.1128/JVI.00426-18. Print 2018 Nov 1. J Virol. 2018. PMID: 30135121 Free PMC article.
-
Marburg Virus VP30 Is Required for Transcription Initiation at the Glycoprotein Gene.mBio. 2022 Oct 26;13(5):e0224322. doi: 10.1128/mbio.02243-22. Epub 2022 Aug 23. mBio. 2022. PMID: 35997284 Free PMC article.
-
Rescue of recombinant Marburg virus from cDNA is dependent on nucleocapsid protein VP30.J Virol. 2006 Jan;80(2):1038-43. doi: 10.1128/JVI.80.2.1038-1043.2006. J Virol. 2006. PMID: 16379005 Free PMC article.
-
Marburg and Ebola viruses.Adv Virus Res. 1996;47:1-52. doi: 10.1016/s0065-3527(08)60733-2. Adv Virus Res. 1996. PMID: 8895830 Review. No abstract available.
-
Marburg Virus Reverse Genetics Systems.Viruses. 2016 Jun 22;8(6):178. doi: 10.3390/v8060178. Viruses. 2016. PMID: 27338448 Free PMC article. Review.
Cited by
-
Filoviral immune evasion mechanisms.Viruses. 2011 Sep;3(9):1634-49. doi: 10.3390/v3091634. Epub 2011 Sep 7. Viruses. 2011. PMID: 21994800 Free PMC article. Review.
-
Recombinant Marburg virus expressing EGFP allows rapid screening of virus growth and real-time visualization of virus spread.J Infect Dis. 2011 Nov;204 Suppl 3(Suppl 3):S861-70. doi: 10.1093/infdis/jir308. J Infect Dis. 2011. PMID: 21987762 Free PMC article.
-
Marburg virus evades interferon responses by a mechanism distinct from ebola virus.PLoS Pathog. 2010 Jan 15;6(1):e1000721. doi: 10.1371/journal.ppat.1000721. PLoS Pathog. 2010. PMID: 20084112 Free PMC article.
-
Virus nomenclature below the species level: a standardized nomenclature for filovirus strains and variants rescued from cDNA.Arch Virol. 2014 May;159(5):1229-37. doi: 10.1007/s00705-013-1877-2. Epub 2013 Nov 5. Arch Virol. 2014. PMID: 24190508 Free PMC article.
-
Minigenomes, transcription and replication competent virus-like particles and beyond: reverse genetics systems for filoviruses and other negative stranded hemorrhagic fever viruses.Antiviral Res. 2011 Aug;91(2):195-208. doi: 10.1016/j.antiviral.2011.06.003. Epub 2011 Jun 14. Antiviral Res. 2011. PMID: 21699921 Free PMC article. Review.
References
-
- Buchholz, U. J., S. Biacchesi, Q. N. Pham, K. C. Tran, L. Yang, C. L. Luongo, M. H. Skiadopoulos, B. R. Murphy, and P. L. Collins. 2005. Deletion of M2 gene open reading frames 1 and 2 of human metapneumovirus: effects on RNA synthesis, attenuation, and immunogenicity. J. Virol. 796588-6597. - PMC - PubMed
-
- Calain, P., M. C. Monroe, and S. T. Nichol. 1999. Ebola virus defective interfering particles and persistent infection. Virology 262114-128. - PubMed
-
- Collins, P. L., E. Camargo, and M. G. Hill. 1999. Support plasmids and support proteins required for recovery of recombinant respiratory syncytial virus. Virology 259251-255. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Medical

