Genome-wide control of the distribution of meiotic recombination
- PMID: 19226188
- PMCID: PMC2642883
- DOI: 10.1371/journal.pbio.1000035
Genome-wide control of the distribution of meiotic recombination
Abstract
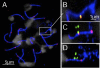
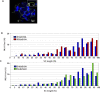
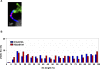
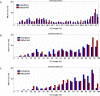
Meiotic recombination events are not randomly distributed in the genome but occur in specific regions called recombination hotspots. Hotspots are predicted to be preferred sites for the initiation of meiotic recombination and their positions and activities are regulated by yet-unknown controls. The activity of the Psmb9 hotspot on mouse Chromosome 17 (Chr 17) varies according to genetic background. It is active in strains carrying a recombinant Chr 17 where the proximal third is derived from Mus musculus molossinus. We have identified the genetic locus required for Psmb9 activity, named Dsbc1 for Double-strand break control 1, and mapped this locus within a 6.7-Mb region on Chr 17. Based on cytological analysis of meiotic DNA double-strand breaks (DSB) and crossovers (COs), we show that Dsbc1 influences DSB and CO, not only at Psmb9, but in several other regions of Chr 17. We further show that CO distribution is also influenced by Dsbc1 on Chrs 15 and 18. Finally, we provide direct molecular evidence for the regulation in trans mediated by Dsbc1, by showing that it controls the CO activity at the Hlx1 hotspot on Chr 1. We thus propose that Dsbc1 encodes for a trans-acting factor involved in the specification of initiation sites of meiotic recombination genome wide in mice.
Conflict of interest statement
Competing interests. The authors have declared that no competing interests exist.
Figures



References
-
- Petronczki M, Siomos MF, Nasmyth K. Un menage a quatre. The molecular biology of chromosome segregation in meiosis. Cell. 2003;112:423–440. - PubMed
-
- Marais G, Charlesworth B. Genome evolution: recombination speeds up adaptive evolution. Curr Biol. 2003;13:R68–70. - PubMed
-
- Hunter N. Meiotic recombination. In: Aguilera A, Rothstein R, editors. Molecular genetics of recombination. Berlin: Springer-Verlag; 2007. pp. 381–442.
-
- Keeney S. Mechanism and control of meiotic recombination initiation. Curr Top Dev Biol. 2001;52:1–53. - PubMed
-
- Baudat F, de Massy B. Regulating double-stranded DNA break repair towards crossover or non-crossover during mammalian meiosis. Chromosome Res. 2007;15:565–577. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Molecular Biology Databases
Miscellaneous

