Parkin, PINK1, and DJ-1 form a ubiquitin E3 ligase complex promoting unfolded protein degradation
- PMID: 19229105
- PMCID: PMC2648688
- DOI: 10.1172/JCI37617
Parkin, PINK1, and DJ-1 form a ubiquitin E3 ligase complex promoting unfolded protein degradation
Abstract
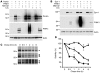
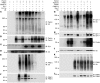
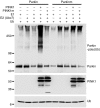
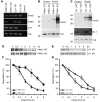
Mutations in PARKIN, pten-induced putative kinase 1 (PINK1), and DJ-1 are individually linked to autosomal recessive early-onset familial forms of Parkinson disease (PD). Although mutations in these genes lead to the same disease state, the functional relationships between them and how their respective disease-associated mutations cause PD are largely unknown. Here, we show that Parkin, PINK1, and DJ-1 formed a complex (termed PPD complex) to promote ubiquitination and degradation of Parkin substrates, including Parkin itself and Synphilin-1 in neuroblastoma cells and human brain lysates. Genetic ablation of either Pink1 or Dj-1 resulted in reduced ubiquitination of endogenous Parkin as well as decreased degradation and increased accumulation of aberrantly expressed Parkin substrates. Expression of PINK1 enhanced Parkin-mediated degradation of heat shock-induced misfolded protein. In contrast, PD-pathogenic Parkin and PINK1 mutations showed reduced ability to promote degradation of Parkin substrates. This study identified a functional ubiquitin E3 ligase complex consisting of PD-associated Parkin, PINK1, and DJ-1 to promote degradation of un-/misfolded proteins and suggests that their PD-pathogenic mutations impair E3 ligase activity of the complex, which may constitute a mechanism underlying PD pathogenesis.
Figures





Comment in
-
Protein degradation in Parkinson disease revisited: it's complex.J Clin Invest. 2009 Mar;119(3):442-5. doi: 10.1172/jci38619. J Clin Invest. 2009. PMID: 19306499 Free PMC article.
References
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Medical
Molecular Biology Databases
Research Materials
Miscellaneous

