Genome stability roles of SUMO-targeted ubiquitin ligases
- PMID: 19233742
- PMCID: PMC2685196
- DOI: 10.1016/j.dnarep.2009.01.010
Genome stability roles of SUMO-targeted ubiquitin ligases
Abstract
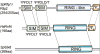
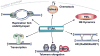
Post-translational modification of the cell's proteome by ubiquitin and ubiquitin-like proteins provides dynamic functional regulation. Ubiquitin and SUMO are well-studied post-translational modifiers that typically impart distinct effects on their targets. The recent discovery that modification by SUMO can target proteins for ubiquitination and proteasomal degradation sets a new paradigm in the field, and offers insights into the roles of SUMO and ubiquitin in genome stability.
Figures

Similar articles
-
Preface. Ubiquitin, SUMO and the maintenance of genome stability.DNA Repair (Amst). 2009 Apr 5;8(4):429. doi: 10.1016/j.dnarep.2009.01.012. Epub 2009 Feb 14. DNA Repair (Amst). 2009. PMID: 19223248 No abstract available.
-
The role of Schizosaccharomyces pombe SUMO ligases in genome stability.Biochem Soc Trans. 2007 Dec;35(Pt 6):1379-84. doi: 10.1042/BST0351379. Biochem Soc Trans. 2007. PMID: 18031226 Review.
-
Mechanism and function of DNA replication-independent DNA-protein crosslink repair via the SUMO-RNF4 pathway.EMBO J. 2021 Sep 15;40(18):e107413. doi: 10.15252/embj.2020107413. Epub 2021 Aug 4. EMBO J. 2021. PMID: 34346517 Free PMC article.
-
SUMO-Targeted Ubiquitin Ligases and Their Functions in Maintaining Genome Stability.Int J Mol Sci. 2021 May 20;22(10):5391. doi: 10.3390/ijms22105391. Int J Mol Sci. 2021. PMID: 34065507 Free PMC article. Review.
-
[DNA damage tolerance through replication bypass of stalled replication forks].Tanpakushitsu Kakusan Koso. 2009 Mar;54(4 Suppl):388-93. Tanpakushitsu Kakusan Koso. 2009. PMID: 21089480 Review. Japanese. No abstract available.
Cited by
-
Role of SUMOylation in differential ERα transcriptional repression by tamoxifen and fulvestrant in breast cancer cells.Oncogene. 2019 Feb;38(7):1019-1037. doi: 10.1038/s41388-018-0468-9. Epub 2018 Sep 6. Oncogene. 2019. PMID: 30190545 Free PMC article.
-
Exploring the RING-catalyzed ubiquitin transfer mechanism by MD and QM/MM calculations.PLoS One. 2014 Jul 8;9(7):e101663. doi: 10.1371/journal.pone.0101663. eCollection 2014. PLoS One. 2014. PMID: 25003393 Free PMC article.
-
Changing the ubiquitin landscape during viral manipulation of the DNA damage response.FEBS Lett. 2011 Sep 16;585(18):2897-906. doi: 10.1016/j.febslet.2011.04.049. Epub 2011 May 5. FEBS Lett. 2011. PMID: 21549706 Free PMC article. Review.
-
SUMOylation inhibits FOXM1 activity and delays mitotic transition.Oncogene. 2014 Aug 21;33(34):4316-29. doi: 10.1038/onc.2013.546. Epub 2013 Dec 23. Oncogene. 2014. PMID: 24362530 Free PMC article.
-
DNA repair and global sumoylation are regulated by distinct Ubc9 noncovalent complexes.Mol Cell Biol. 2011 Jun;31(11):2299-310. doi: 10.1128/MCB.05188-11. Epub 2011 Mar 28. Mol Cell Biol. 2011. PMID: 21444718 Free PMC article.
References
-
- Lallemand-Breitenbach V, Zhu J, Puvion F, Koken M, Honore N, Doubeikovsky A, Duprez E, Pandolfi PP, Puvion E, Freemont P, et al. Role of promyelocytic leukemia (PML) sumolation in nuclear body formation, 11S proteasome recruitment, and As2O3-induced PML or PML/retinoic acid receptor alpha degradation. J Exp Med. 2001;193:1361–1371. - PMC - PubMed
-
- Perry JJ, Tainer JA, Boddy MN. A SIM-ultaneous role for SUMO and ubiquitin. Trends Biochem Sci. 2008;33:201–208. - PubMed
-
- Lallemand-Breitenbach V, Jeanne M, Benhenda S, Nasr R, Lei M, Peres L, Zhou J, Zhu J, Raught B, de The H. Arsenic degrades PML or PML-RARalpha through a SUMO-triggered RNF4/ubiquitin-mediated pathway. Nat Cell Biol. 2008;10:547–555. - PubMed
-
- Tatham MH, Geoffroy MC, Shen L, Plechanovova A, Hattersley N, Jaffray EG, Palvimo JJ, Hay RT. RNF4 is a poly-SUMO-specific E3 ubiquitin ligase required for arsenic-induced PML degradation. Nat Cell Biol. 2008;10:538–546. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources

