Combined analysis of transcriptome and proteome data as a tool for the identification of candidate biomarkers in renal cell carcinoma
- PMID: 19235166
- PMCID: PMC3410642
- DOI: 10.1002/pmic.200700288
Combined analysis of transcriptome and proteome data as a tool for the identification of candidate biomarkers in renal cell carcinoma
Abstract
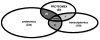
Results obtained from expression profilings of renal cell carcinoma using different "ome"-based approaches and comprehensive data analysis demonstrated that proteome-based technologies and cDNA microarray analyses complement each other during the discovery phase for disease-related candidate biomarkers. The integration of the respective data revealed the uniqueness and complementarities of the different technologies. While comparative cDNA microarray analyses though restricted to up-regulated targets largely revealed genes involved in controlling gene/protein expression (19%) and signal transduction processes (13%), proteomics/PROTEOMEX-defined candidate biomarkers include enzymes of the cellular metabolism (36%), transport proteins (12%), and cell motility/structural molecules (10%). Candidate biomarkers defined by proteomics and PROTEOMEX are frequently shared, whereas the sharing rate between cDNA microarray and proteome-based profilings is limited. Putative candidate biomarkers provide insights into their cellular (dys)function and their diagnostic/prognostic value but still warrant further validation in larger patient numbers. Based on the fact that merely three candidate biomarkers were shared by all applied technologies, namely annexin A4, tubulin alpha-1A chain, and ubiquitin carboxyl-terminal hydrolase L1, the analysis at a single hierarchical level of biological regulation seems to provide only limited results thus emphasizing the importance and benefit of performing rather combinatorial screenings which can complement the standard clinical predictors.
Figures



References
-
- Ding C, Cantor CR. Quantitative analysis of nucleic acids--the last few years of progress. J Biochem Mol Biol. 2004;37:1–10. - PubMed
-
- Guo QM. DNA microarray and cancer. Curr Opin Oncol. 2003;15:36–43. - PubMed
-
- Waters KM, Pounds JG, Thrall BD. Data merging for integrated microarray and proteomic analysis. Brief Funct Genomic Proteomic. 2006;5:261–272. - PubMed
-
- Tian Q, Stepaniants SB, Mao M, Weng L, et al. Integrated genomic and proteomic analyses of gene expression in Mammalian cells. Mol Cell Proteomics. 2004;3:960–969. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Medical

