Substrate binding in the FAD-dependent hydroxynitrile lyase from almond provides insight into the mechanism of cyanohydrin formation and explains the absence of dehydrogenation activity
- PMID: 19256550
- PMCID: PMC2669238
- DOI: 10.1021/bi802162s
Substrate binding in the FAD-dependent hydroxynitrile lyase from almond provides insight into the mechanism of cyanohydrin formation and explains the absence of dehydrogenation activity
Abstract
In a large number of plant species hydroxynitrile lyases catalyze the decomposition of cyanohydrins in order to generate hydrogen cyanide upon tissue damage. Hydrogen cyanide serves as a deterrent against herbivores and fungi. In vitro hydroxynitrile lyases are proficient biocatalysts for the stereospecific synthesis of cyanohydrins. Curiously, hydroxynitrile lyases from different species are completely unrelated in structure and substrate specificity despite catalyzing the same reaction. The hydroxynitrile lyase from almond shows close resemblance to flavoproteins of the glucose-methanol-choline oxidoreductase family. We report here 3D structural data of this lyase with the reaction product benzaldehyde bound within the active site, which allow unambiguous assignment of the location of substrate binding. Based on the binding geometry, a reaction mechanism is proposed that involves one of the two conserved active site histidine residues acting as a general base abstracting the proton from the cyanohydrin hydroxyl group. Site-directed mutagenesis shows that both active site histidines are required for the reaction to occur. There is no evidence that the flavin cofactor directly participates in the reaction. Comparison with other hydroxynitrile lyases reveals a large diversity of active site architectures, which, however, share the common features of a general active site base and a nearby patch with positive electrostatic potential. On the basis of the difference in substrate binding geometry between the FAD-dependent HNL from almond and the related oxidases, we can rationalize why the HNL does not act as an oxidase.
Figures


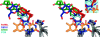

References
-
- Conn E. E. (1981) in The Biochemistry of Plants: A Comprehensive Treatise (Stumpf P. K., and Conn E. E., Eds.) pp 479−500, Academic Press, New York.
-
- Nahrstedt A. (1985) Cyanogenic compounds as protecting agents for organisms. Plant Syst. Evol. 150, 35–47.
-
- Hickel A.; Hasslacher M.; Griengl H. (1996) Hydroxynitrile lyases: functions and properties. Physiol. Plant. 98, 891–898.
-
- Wajant H.; Effenberger F. (1996) Hydroxynitrile lyases of higher plants. Biol. Chem. 377, 611–617. - PubMed
-
- Purkarthofer T.; Skranc W.; Schuster C.; Griengl H. (2007) Potential and capabilities of hydroxynitrile lyases as biocatalysts in the chemical industry. Appl. Microbiol. Biotechnol. 76, 309–320. - PubMed
Publication types
MeSH terms
Substances
Associated data
- Actions
- Actions
Grants and funding
LinkOut - more resources
Full Text Sources
Molecular Biology Databases
Miscellaneous

