Fractured genes: a novel genomic arrangement involving new split inteins and a new homing endonuclease family
- PMID: 19264795
- PMCID: PMC2677866
- DOI: 10.1093/nar/gkp095
Fractured genes: a novel genomic arrangement involving new split inteins and a new homing endonuclease family
Abstract
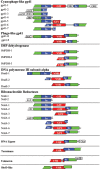
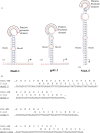
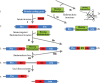
Inteins are genetic elements, inserted in-frame into protein-coding genes, whose products catalyze their removal from the protein precursor via a protein-splicing reaction. Intein domains can be split into two fragments and still ligate their flanks by a trans-protein-splicing reaction. A bioinformatic analysis of environmental metagenomic data revealed 26 different loci with a novel genomic arrangement. In each locus, a conserved enzyme coding region is broken in two by a split intein, with a free-standing endonuclease gene inserted in between. Eight types of DNA synthesis and repair enzymes have this 'fractured' organization. The new types of naturally split-inteins were analyzed in comparison to known split-inteins. Some loci include apparent gene control elements brought in with the endonuclease gene. A newly predicted homing endonuclease family, related to very-short patch repair (Vsr) endonucleases, was found in half of the loci. These putative homing endonucleases also appear in group-I introns, and as stand-alone inserts in the absence of surrounding intervening sequences. The new fractured genes organization appears to be present mainly in phage, shows how endonucleases can integrate into inteins, and may represent a missing link in the evolution of gene breaking in general, and in the creation of split-inteins in particular.
Figures


Similar articles
-
Genetic definition of a protein-splicing domain: functional mini-inteins support structure predictions and a model for intein evolution.Proc Natl Acad Sci U S A. 1997 Oct 14;94(21):11466-71. doi: 10.1073/pnas.94.21.11466. Proc Natl Acad Sci U S A. 1997. PMID: 9326633 Free PMC article.
-
Modular organization of inteins and C-terminal autocatalytic domains.Protein Sci. 1998 Jan;7(1):64-71. doi: 10.1002/pro.5560070106. Protein Sci. 1998. PMID: 9514260 Free PMC article.
-
Homing endonucleases residing within inteins: evolutionary puzzles awaiting genetic solutions.Biochem Soc Trans. 2011 Jan;39(1):169-73. doi: 10.1042/BST0390169. Biochem Soc Trans. 2011. PMID: 21265767 Review.
-
Conserved sequence features of inteins (protein introns) and their use in identifying new inteins and related proteins.Protein Sci. 1994 Dec;3(12):2340-50. doi: 10.1002/pro.5560031218. Protein Sci. 1994. PMID: 7756989 Free PMC article.
-
Structural and functional characteristics of homing endonucleases.Crit Rev Biochem Mol Biol. 2003;38(3):199-248. doi: 10.1080/713609235. Crit Rev Biochem Mol Biol. 2003. PMID: 12870715 Review.
Cited by
-
A cybergenetic framework for engineering intein-mediated integral feedback control systems.Nat Commun. 2023 Mar 11;14(1):1337. doi: 10.1038/s41467-023-36863-9. Nat Commun. 2023. PMID: 36906662 Free PMC article.
-
Nuclease genes occupy boundaries of genetic exchange between bacteriophages.NAR Genom Bioinform. 2023 Aug 24;5(3):lqad076. doi: 10.1093/nargab/lqad076. eCollection 2023 Sep. NAR Genom Bioinform. 2023. PMID: 37636022 Free PMC article.
-
Homing endonucleases: from genetic anomalies to programmable genomic clippers.Methods Mol Biol. 2014;1123:1-26. doi: 10.1007/978-1-62703-968-0_1. Methods Mol Biol. 2014. PMID: 24510256 Free PMC article. Review.
-
Model-guided design of mammalian genetic programs.Sci Adv. 2021 Feb 19;7(8):eabe9375. doi: 10.1126/sciadv.abe9375. Print 2021 Feb. Sci Adv. 2021. PMID: 33608279 Free PMC article.
-
Near-Infrared Optogenetic Module for Conditional Protein Splicing.J Mol Biol. 2023 Dec 15;435(24):168360. doi: 10.1016/j.jmb.2023.168360. Epub 2023 Nov 8. J Mol Biol. 2023. PMID: 37949312 Free PMC article.
References
-
- Haugen P, Simon DM, Bhattacharya D. The natural history of group I introns. Trends Genet. 2005;21:111–119. - PubMed
-
- Dassa B, Haviv H, Amitai G, Pietrokovski S. Protein splicing and auto-cleavage of bacterial intein-like domains lacking a C′-flanking nucleophilic residue. J. Biol. Chem. 2004;279:32001–32007. - PubMed
-
- Dassa B, Yanai I, Pietrokovski S. New type of polyubiquitin-like genes with intein-like autoprocessing domains. Trends Genet. 2004;20:538–542. - PubMed
-
- Marcotte EM, Pellegrini M, Ng HL, Rice DW, Yeates TO, Eisenberg D. Detecting protein function and protein-protein interactions from genome sequences. Science. 1999;285:751–753. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources

