Threshold occupancy and specific cation binding modes in the hammerhead ribozyme active site are required for active conformation
- PMID: 19265710
- PMCID: PMC2715853
- DOI: 10.1016/j.jmb.2009.02.054
Threshold occupancy and specific cation binding modes in the hammerhead ribozyme active site are required for active conformation
Abstract
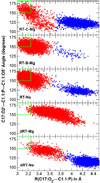
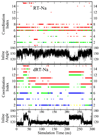
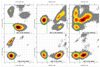
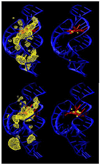
The relationship between formation of active in-line attack conformations and monovalent (Na(+)) and divalent (Mg(2+)) metal ion binding in hammerhead ribozyme (HHR) has been explored with molecular dynamics simulations. To stabilize repulsions between negatively charged groups, different requirements of the threshold occupancy of metal ions were observed in the reactant and activated precursor states both in the presence and in the absence of a Mg(2+) in the active site. Specific bridging coordination patterns of the ions are correlated with the formation of active in-line attack conformations and can be accommodated in both cases. Furthermore, simulation results suggest that the HHR folds to form an electronegative recruiting pocket that attracts high local concentrations of positive charge. The present simulations help to reconcile experiments that probe the metal ion sensitivity of HHR catalysis and support the supposition that Mg(2+), in addition to stabilizing active conformations, plays a specific chemical role in catalysis.
Figures

Similar articles
-
Metal binding motif in the active site of the HDV ribozyme binds divalent and monovalent ions.Biochemistry. 2011 Apr 5;50(13):2672-82. doi: 10.1021/bi2000164. Epub 2011 Mar 9. Biochemistry. 2011. PMID: 21348498 Free PMC article.
-
Bridging the gap between theory and experiment to derive a detailed understanding of hammerhead ribozyme catalysis.Prog Mol Biol Transl Sci. 2013;120:25-91. doi: 10.1016/B978-0-12-381286-5.00002-0. Prog Mol Biol Transl Sci. 2013. PMID: 24156941 Free PMC article. Review.
-
Role of Mg2+ in hammerhead ribozyme catalysis from molecular simulation.J Am Chem Soc. 2008 Mar 12;130(10):3053-64. doi: 10.1021/ja076529e. Epub 2008 Feb 14. J Am Chem Soc. 2008. PMID: 18271579 Free PMC article.
-
Existence of efficient divalent metal ion-catalyzed and inefficient divalent metal ion-independent channels in reactions catalyzed by a hammerhead ribozyme.Nucleic Acids Res. 2002 Jun 1;30(11):2374-82. doi: 10.1093/nar/30.11.2374. Nucleic Acids Res. 2002. PMID: 12034824 Free PMC article.
-
Two distinct catalytic strategies in the hepatitis δ virus ribozyme cleavage reaction.Biochemistry. 2011 Nov 8;50(44):9424-33. doi: 10.1021/bi201157t. Epub 2011 Oct 17. Biochemistry. 2011. PMID: 22003985 Free PMC article. Review.
Cited by
-
Bulky cations greatly increase the turnover of a native hammerhead ribozyme.RSC Adv. 2019 Nov 4;9(61):35820-35824. doi: 10.1039/c9ra06797c. eCollection 2019 Oct 31. RSC Adv. 2019. PMID: 35528091 Free PMC article.
-
Dynamical ensemble of the active state and transition state mimic for the RNA-cleaving 8-17 DNAzyme in solution.Nucleic Acids Res. 2019 Nov 4;47(19):10282-10295. doi: 10.1093/nar/gkz773. Nucleic Acids Res. 2019. PMID: 31511899 Free PMC article.
-
Divalent Metal Ion Activation of a Guanine General Base in the Hammerhead Ribozyme: Insights from Molecular Simulations.Biochemistry. 2017 Jun 20;56(24):2985-2994. doi: 10.1021/acs.biochem.6b01192. Epub 2017 Jun 12. Biochemistry. 2017. PMID: 28530384 Free PMC article.
-
Metal binding motif in the active site of the HDV ribozyme binds divalent and monovalent ions.Biochemistry. 2011 Apr 5;50(13):2672-82. doi: 10.1021/bi2000164. Epub 2011 Mar 9. Biochemistry. 2011. PMID: 21348498 Free PMC article.
-
Ribozyme Catalysis with a Twist: Active State of the Twister Ribozyme in Solution Predicted from Molecular Simulation.J Am Chem Soc. 2016 Mar 9;138(9):3058-65. doi: 10.1021/jacs.5b12061. Epub 2016 Feb 25. J Am Chem Soc. 2016. PMID: 26859432 Free PMC article.
References
-
- Scott WG. Ribozymes. Curr. Opin. Struct. Biol. 2007;13:280–286. - PubMed
-
- Scott WG. Biophysical and biochemical investigations of RNA catalysis in the hammerhead ribozyme. Q. Rev. Biophys. 1999;32:241–294. - PubMed
-
- Scott WG. Morphing the minimal and full-length hammerhead ribozymes: implications for the cleavage mechanism. Biol. Chem. 2007;388:727–735. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources

