Genetic variation and population substructure in outbred CD-1 mice: implications for genome-wide association studies
- PMID: 19266100
- PMCID: PMC2649211
- DOI: 10.1371/journal.pone.0004729
Genetic variation and population substructure in outbred CD-1 mice: implications for genome-wide association studies
Abstract
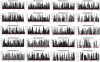
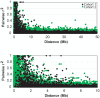
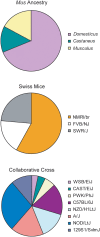
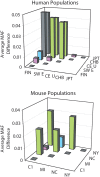
Outbred laboratory mouse populations are widely used in biomedical research. Since little is known about the degree of genetic variation present in these populations, they are not widely used for genetic studies. Commercially available outbred CD-1 mice are drawn from an extremely large breeding population that has accumulated many recombination events, which is desirable for genome-wide association studies. We therefore examined the degree of genome-wide variation within CD-1 mice to investigate their suitability for genetic studies. The CD-1 mouse genome displays patterns of linkage disequilibrium and heterogeneity similar to wild-caught mice. Population substructure and phenotypic differences were observed among CD-1 mice obtained from different breeding facilities. Differences in genetic variation among CD-1 mice from distinct facilities were similar to genetic differences detected between closely related human populations, consistent with a founder effect. This first large-scale genetic analysis of the outbred CD-1 mouse strain provides important considerations for the design and analysis of genetic studies in CD-1 mice.
Conflict of interest statement
Figures
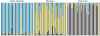

References
-
- Cui S, Chesson C, Hope R. Genetic variation within and between strains of outbred Swiss mice. Lab Anim. 1993;27:116–123. - PubMed
-
- Rice MC, O'Brien SJ. Genetic variance of laboratory outbred Swiss mice. Nature. 1980;283:157–161. - PubMed
-
- Chia R, Achilli F, Festing MF, Fisher EM. The origins and uses of mouse outbred stocks. Nat Genet. 2005;37:1181–1186. - PubMed
-
- Brown JA, Chua SC, Jr, Liu SM, Andrews MT, Vandenbergh JG. Spontaneous mutation in the db gene results in obesity and diabetes in CD-1 outbred mice. Am J Physiol Regul Integr Comp Physiol. 2000;278:R320–330. - PubMed
-
- Lehoczky JA, Cai WW, Douglas JA, Moran JL, Beier DR, et al. Description and genetic mapping of Polypodia: an X-linked dominant mouse mutant with ectopic caudal limbs and other malformations. Mamm Genome. 2006;17:903–913. - PubMed
Publication types
MeSH terms
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases

