Spt6 enhances the elongation rate of RNA polymerase II in vivo
- PMID: 19279664
- PMCID: PMC2683705
- DOI: 10.1038/emboj.2009.56
Spt6 enhances the elongation rate of RNA polymerase II in vivo
Abstract
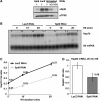
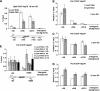
Several eukaryotic transcription factors have been shown to modulate the elongation rate of RNA polymerase II (Pol II) on naked or chromatin-reconstituted templates in vitro. However, none of the tested factors have been shown to directly affect the elongation rate of Pol II in vivo. We performed a directed RNAi knock-down (KD) screen targeting 141 candidate transcription factors and identified multiple factors, including Spt6, that alter the induced Hsp70 transcript levels in Drosophila S2 cells. Spt6 is known to interact with both nucleosome structure and Pol II, and it has properties consistent with having a role in elongation. Here, ChIP assays of the first wave of Pol II after heat shock in S2 cells show that KD of Spt6 reduces the rate of Pol II elongation. Also, fluorescence recovery after photobleaching assays of GFP-Pol II in salivary gland cells show that this Spt6-dependent effect on elongation rate persists during steady-state-induced transcription, reducing the elongation rate from approximately 1100 to 500 bp/min. Furthermore, RNAi depletion of Spt6 reveals its broad requirement during different stages of development.
Figures




References
-
- Adelman K, Marr MT, Werner J, Saunders A, Ni Z, Andrulis ED, Lis JT (2005) Efficient release from promoter-proximal stall sites requires transcript cleavage factor TFIIS. Mol Cell 17: 103–112 - PubMed
-
- Adkins MW, Tyler JK (2006) Transcriptional activators are dispensable for transcription in the absence of Spt6-mediated chromatin reassembly of promoter regions. Mol Cell 21: 405–416 - PubMed
-
- Andrulis ED, Werner J, Nazarian A, Erdjument-Bromage H, Tempst P, Lis JT (2002) The RNA processing exosome is linked to elongating RNA polymerase II in Drosophila. Nature 420: 837–841 - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Molecular Biology Databases

