Bacterial cell curvature through mechanical control of cell growth
- PMID: 19279668
- PMCID: PMC2683044
- DOI: 10.1038/emboj.2009.61
Bacterial cell curvature through mechanical control of cell growth
Abstract
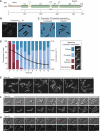
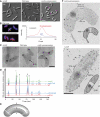
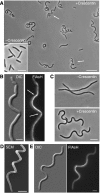
The cytoskeleton is a key regulator of cell morphogenesis. Crescentin, a bacterial intermediate filament-like protein, is required for the curved shape of Caulobacter crescentus and localizes to the inner cell curvature. Here, we show that crescentin forms a single filamentous structure that collapses into a helix when detached from the cell membrane, suggesting that it is normally maintained in a stretched configuration. Crescentin causes an elongation rate gradient around the circumference of the sidewall, creating a longitudinal cell length differential and hence curvature. Such curvature can be produced by physical force alone when cells are grown in circular microchambers. Production of crescentin in Escherichia coli is sufficient to generate cell curvature. Our data argue for a model in which physical strain borne by the crescentin structure anisotropically alters the kinetics of cell wall insertion to produce curved growth. Our study suggests that bacteria may use the cytoskeleton for mechanical control of growth to alter morphology.
Figures




Comment in
-
Bend into shape.EMBO J. 2009 May 6;28(9):1193-4. doi: 10.1038/emboj.2009.91. EMBO J. 2009. PMID: 19421162 Free PMC article. No abstract available.
References
-
- Aaron M, Charbon G, Lam H, Schwarz H, Vollmer W, Jacobs-Wagner C (2007) The tubulin homologue FtsZ contributes to cell elongation by guiding cell wall precursor synthesis in Caulobacter crescentus. Mol Microbiol 64: 938–952 - PubMed
-
- Arnoldi M, Fritz M, Bauerlein E, Radmacher M, Sackmann E, Boulbitch A (2000) Bacterial turgor pressure can be measured by atomic force microscopy. Phys Rev E Stat Phys Plasmas Fluids Relat Interdiscip Topics 62 (1 Part B): 1034–1044 - PubMed
-
- Ausmees N, Kuhn JR, Jacobs-Wagner C (2003) The bacterial cytoskeleton: an intermediate filament-like function in cell shape. Cell 115: 705–713 - PubMed
-
- Bertsche U, Kast T, Wolf B, Fraipont C, Aarsman ME, Kannenberg K, von Rechenberg M, Nguyen-Disteche M, den Blaauwen T, Höltje JV, Vollmer W (2006) Interaction between two murein (peptidoglycan) synthases, PBP3 and PBP1B, in Escherichia coli. Mol Microbiol 61: 675–690 - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Molecular Biology Databases

