A theoretical model for the mechanical unfolding of repeat proteins
- PMID: 19289042
- PMCID: PMC2717355
- DOI: 10.1016/j.bpj.2008.12.3899
A theoretical model for the mechanical unfolding of repeat proteins
Abstract
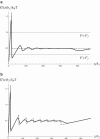
We consider the mechanical stretching of a polypeptide chain formed by multiple interacting repeats. The folding thermodynamics and the interactions among the repeats are described by the Ising model. Unfolded repeats act as soft entropic springs, whereas folded repeats respond to a force as stiffer springs. We show that the resulting force-extension curve may exhibit a pronounced force maximum corresponding to the unfolding of the first repeat. This event is followed by the unfolding of the remaining repeats, which takes place at a lower force. As the protein extension is increased, the force-extension curve of a sufficiently long repeat protein displays a plateau, where the force remains nearly constant and the protein unfolds sequentially so that the number of unfolded repeats is proportional to the extension. Such a sequential mechanical unfolding mechanism is displayed even by the repeat proteins whose thermal denaturation is highly cooperative, provided that they are long enough. By contrast, the unfolding of short repeat progressions can be cooperative.
Figures
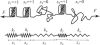

References
-
- Kajander T., Cortajarena A.L., Main E.R., Mochrie S.G., Regan L. A new folding paradigm for repeat proteins. J. Am. Chem. Soc. 2005;127:10188–10190. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Research Materials

