The role of genomic data in the discovery, annotation and evolutionary interpretation of the interferon-lambda family
- PMID: 19300512
- PMCID: PMC2654155
- DOI: 10.1371/journal.pone.0004933
The role of genomic data in the discovery, annotation and evolutionary interpretation of the interferon-lambda family
Abstract
Background: Type-I interferons, type-II interferons, and the IL-10 family are helical cytokines with similar three-dimensional folds. However, their homologous relationship is difficult to detect on the basis of sequence alone. We have previously described the discovery of the human type-III interferons (IFN lambda-1, -2, -3 or IL-29, IL-28A, IL-28B), which required a combination of manual and computational techniques applied to predicted protein sequences.
Principal findings: Here we describe how the use of gene structure analysis and comparative genomics enabled a more extensive understanding of these genes early in the discovery process. More recently, additional mammalian genome sequences have shown that there are between one and potentially nine copies of interferon lambda genes in each genome, and that several species have single exon versions of the interferon lambda gene.
Significance: The variable number of single exon type-I interferons in mammals, along with recently identified genes in zebrafish homologous to interferons allows a story of interferon evolution to be proposed. This model suggests that the gene duplications and single exon retrotransposons of mammalian type-III interferons are positively selected for within a genome. These characteristics are also shared with the fish interferons and could be responsible for the generation of the IL10 family and also the single exon type-I interferons.
Conflict of interest statement
Figures
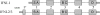

References
-
- Altschul S, Madden T, Schaffer A, Zhang J, Zhang Z, et al. Gapped BLAST and PSI-BLAST: a new generation of protein database search programs. Nucleic Acids Res. 1997;25:3389–3402. http://dx.doi.org/10.1093/nar/25.17.3389. - DOI - PMC - PubMed
-
- Hill E, Morea V, Chothia C. Sequence conservation in families whose members have little or no sequence similarity: the four-helical cytokines and cytochromes. J Mol Biol. 2002;322:205–233. http://view.ncbi.nlm.nih.gov/pubmed/12215425. - PubMed
-
- Blumberg H, Conklin D, Xu W, Grossmann A, Brender T, et al. Interleukin 20: discovery, receptor identification, and role in epidermal function. Cell. 2001;104:9–19. http://view.ncbi.nlm.nih.gov/pubmed/11163236. - PubMed
-
- Conklin D. Recognition of the helical cytokine fold. Journal of computational biology : a journal of computational molecular cell biology. 2004;11:1189–1200. http://dx.doi.org/10.1089/cmb.2004.11.1189. - DOI - PubMed
-
- Conklin D, Haldeman B, Gao Z. Gene finding for the helical cytokines. Bioinformatics. 2005;21 http://dx.doi.org/doi:10.1093/bioinformatics/bti283. - DOI - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Molecular Biology Databases

