PLRG1 is an essential regulator of cell proliferation and apoptosis during vertebrate development and tissue homeostasis
- PMID: 19307306
- PMCID: PMC2682009
- DOI: 10.1128/MCB.01807-08
PLRG1 is an essential regulator of cell proliferation and apoptosis during vertebrate development and tissue homeostasis
Abstract
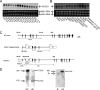
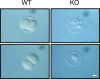
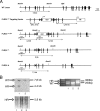
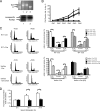
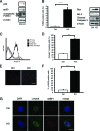
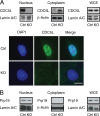
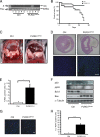
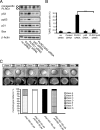
PLRG1, an evolutionarily conserved component of the spliceosome, forms a complex with Pso4/SNEV/Prp19 and the cell division and cycle 5 homolog (CDC5L) that is involved in both pre-mRNA splicing and DNA repair. Here, we show that the inactivation of PLRG1 in mice results in embryonic lethality at 1.5 days postfertilization. Studies of heart- and neuron-specific PLRG1 knockout mice further reveal an essential role of PLRG1 in adult tissue homeostasis and the suppression of apoptosis. PLRG1-deficient mouse embryonic fibroblasts (MEFs) fail to progress through S phase upon serum stimulation and exhibit increased rates of apoptosis. PLRG1 deficiency causes enhanced p53 phosphorylation and stabilization in the presence of increased gamma-H2AX immunoreactivity as an indicator of an activated DNA damage response. p53 downregulation rescues lethality in both PLRG1-deficient MEFs and zebrafish in vivo, showing that apoptosis resulting from PLRG1 deficiency is p53 dependent. Moreover, the deletion of PLRG1 results in the relocation of its interaction partner CDC5L from the nucleus to the cytoplasm without general alterations in pre-mRNA splicing. Taken together, the results of this study identify PLRG1 as a critical nuclear regulator of p53-dependent cell cycle progression and apoptosis during both embryonic development and adult tissue homeostasis.
Figures

References
-
- Agrelo, R., W. H. Cheng, F. Setien, S. Ropero, J. Espada, M. F. Fraga, M. Herranz, M. F. Paz, M. Sanchez-Cespedes, M. J. Artiga, D. Guerrero, A. Castells, C. von Kobbe, V. A. Bohr, and M. Esteller. 2006. Epigenetic inactivation of the premature aging Werner syndrome gene in human cancer. Proc. Natl. Acad. Sci. USA 1038822-8827. - PMC - PubMed
-
- Ajuh, P., J. Sleeman, J. Chusainow, and A. I. Lamond. 2001. A direct interaction between the carboxyl-terminal region of CDC5L and the WD40 domain of PLRG1 is essential for pre-mRNA splicing. J. Biol. Chem. 27642370-42381. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases
Research Materials
Miscellaneous
