On the chimeric nature, thermophilic origin, and phylogenetic placement of the Thermotogales
- PMID: 19307556
- PMCID: PMC2667022
- DOI: 10.1073/pnas.0901260106
On the chimeric nature, thermophilic origin, and phylogenetic placement of the Thermotogales
Abstract
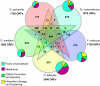
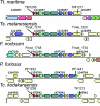
Since publication of the first Thermotogales genome, Thermotoga maritima strain MSB8, single- and multi-gene analyses have disagreed on the phylogenetic position of this order of Bacteria. Here we present the genome sequences of 4 additional members of the Thermotogales (Tt. petrophila, Tt. lettingae, Thermosipho melanesiensis, and Fervidobacterium nodosum) and a comprehensive comparative analysis including the original T. maritima genome. While ribosomal protein genes strongly place Thermotogales as a sister group to Aquificales, the majority of genes with sufficient phylogenetic signal show affinities to Archaea and Firmicutes, especially Clostridia. Indeed, on the basis of the majority of genes in their genomes (including genes that are also found in Aquificales), Thermotogales should be considered members of the Firmicutes. This result highlights the conflict between the taxonomic goal of assigning every species to a unique position in an inclusive Linnaean hierarchy and the evolutionary goal of understanding phylogenesis in the presence of pervasive horizontal gene transfer (HGT) within prokaryotes. Amino acid compositions of reconstructed ancestral sequences from 423 gene families suggest an origin of this gene pool even more thermophilic than extant members of this order, followed by adaptation to lower growth temperatures within the Thermotogales.
Conflict of interest statement
The authors declare no conflict of interest.
Figures


References
-
- Nelson KE, et al. Evidence for lateral gene transfer between Archaea and Bacteria from genome sequence of Thermotoga maritima. Nature. 1999;399:323–329. - PubMed
Publication types
MeSH terms
Associated data
- Actions
- Actions
- Actions
- Actions
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases

