Spatiotemporal profiling of starch biosynthesis and degradation in the developing barley grain
- PMID: 19321714
- PMCID: PMC2675734
- DOI: 10.1104/pp.108.133520
Spatiotemporal profiling of starch biosynthesis and degradation in the developing barley grain
Abstract
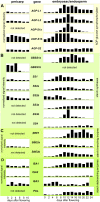
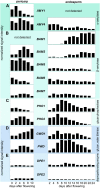
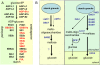
Barley (Hordeum vulgare) grains synthesize starch as the main storage compound. However, some starch is degraded already during caryopsis development. We studied temporal and spatial expression patterns of genes coding for enzymes of starch synthesis and degradation. These profiles coupled with measurements of selected enzyme activities and metabolites have allowed us to propose a role for starch degradation in maternal and filial tissues of developing grains. Early maternal pericarp functions as a major short-term starch storage tissue, possibly ensuring sink strength of the young caryopsis. Gene expression patterns and enzyme activities suggest two different pathways for starch degradation in maternal tissues. One pathway possibly occurs via alpha-amylases 1 and 4 and beta-amylase 1 in pericarp, nucellus, and nucellar projection, tissues that undergo programmed cell death. Another pathway is deducted for living pericarp and chlorenchyma cells, where transient starch breakdown correlates with expression of chloroplast-localized beta-amylases 5, 6, and 7, glucan, water dikinase 1, phosphoglucan, water dikinase, isoamylase 3, and disproportionating enzyme. The suite of genes involved in starch synthesis in filial starchy endosperm is much more complex than in pericarp and involves several endosperm-specific genes. Transient starch turnover occurs in transfer cells, ensuring the maintenance of sink strength in filial tissues and the reallocation of sugars into more proximal regions of the starchy endosperm. Starch is temporally accumulated also in aleurone cells, where it is degraded during the seed filling period, to be replaced by storage proteins and lipids.
Figures







Similar articles
-
The role of invertases and hexose transporters in controlling sugar ratios in maternal and filial tissues of barley caryopses during early development.Plant J. 2003 Jan;33(2):395-411. doi: 10.1046/j.1365-313x.2003.01633.x. Plant J. 2003. PMID: 12535352
-
Identification of genes specifically expressed in maternal and filial tissues of barley caryopses: a cDNA array analysis.Mol Genet Genomics. 2002 Jan;266(5):758-67. doi: 10.1007/s00438-001-0614-9. Epub 2001 Nov 27. Mol Genet Genomics. 2002. PMID: 11810249
-
Isolation of tissues and preservation of RNA from intact, germinated barley grain.Plant J. 2017 Aug;91(4):754-765. doi: 10.1111/tpj.13600. Epub 2017 Jun 15. Plant J. 2017. PMID: 28509349
-
Development of maternal seed tissue in barley is mediated by regulated cell expansion and cell disintegration and coordinated with endosperm growth.J Exp Bot. 2011 Jan;62(3):1217-27. doi: 10.1093/jxb/erq348. Epub 2010 Nov 8. J Exp Bot. 2011. PMID: 21059741 Free PMC article.
-
Barley grain development toward an integrative view.Int Rev Cell Mol Biol. 2010;281:49-89. doi: 10.1016/S1937-6448(10)81002-0. Int Rev Cell Mol Biol. 2010. PMID: 20460183 Review.
Cited by
-
Mapping dynamic QTL dissects the genetic architecture of grain size and grain filling rate at different grain-filling stages in barley.Sci Rep. 2019 Dec 11;9(1):18823. doi: 10.1038/s41598-019-53620-5. Sci Rep. 2019. PMID: 31827117 Free PMC article.
-
Differential expression of two β-amylase genes (Bmy1 and Bmy2) in developing and mature barley grain.Planta. 2011 May;233(5):1001-10. doi: 10.1007/s00425-011-1348-5. Epub 2011 Jan 30. Planta. 2011. PMID: 21279650
-
Rice caryopsis development I: Dynamic changes in different cell layers.J Integr Plant Biol. 2016 Sep;58(9):772-85. doi: 10.1111/jipb.12440. Epub 2015 Dec 16. J Integr Plant Biol. 2016. PMID: 26472484 Free PMC article.
-
A Potential Role of Flag Leaf Potassium in Conferring Tolerance to Drought-Induced Leaf Senescence in Barley.Front Plant Sci. 2016 Feb 26;7:206. doi: 10.3389/fpls.2016.00206. eCollection 2016. Front Plant Sci. 2016. PMID: 26955376 Free PMC article.
-
Barley callus: a model system for bioengineering of starch in cereals.Plant Methods. 2012 Sep 7;8(1):36. doi: 10.1186/1746-4811-8-36. Plant Methods. 2012. PMID: 22958600 Free PMC article.
References
-
- Allison MJ, Ellis RP, Swanston JS (1974) Tissue distribution of α-amylase and phosphorylase in developing barley grains. J Inst Brew 80 488–491
-
- Asatsuma S, Sawada C, Itoh K, Okito M, Kitajima A, Mitsui T (2005) Involvement of α-amylase I-1 in starch degradation in rice chloroplasts. Plant Cell Physiol 46 858–869 - PubMed
-
- Bak-Jensen KS, Laugesen S, Ostergaard O, Finnie C, Roepstorff P, Svensson B (2007) Spatio-temporal profiling and degradation of α-amylase isozymes during barley seed germination. FEBS J 274 2552–2565 - PubMed
-
- Baunsgaard L, Lutken H, Mikkelsen R, Glaring MA, Pham TT, Blennow A (2005) A novel isoform of glucan, water dikinase phosphorylates pre-phosphorylated α-glucans and is involved in starch degradation in Arabidopsis. Plant J 41 595–605 - PubMed
-
- Beck E, Ziegler P (1989) Biosynthesis and degradation of starch in higher plants. Annu Rev Plant Physiol Plant Mol Biol 40 95–117
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources

