Separate domains of Rev1 mediate two modes of DNA damage bypass in mammalian cells
- PMID: 19332561
- PMCID: PMC2682010
- DOI: 10.1128/MCB.00071-09
Separate domains of Rev1 mediate two modes of DNA damage bypass in mammalian cells
Abstract
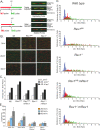
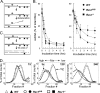
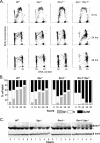
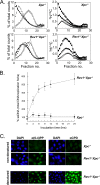
The Y family DNA polymerase Rev1 has been proposed to play a regulatory role in the replication of damaged templates. To elucidate the mechanism by which Rev1 promotes DNA damage bypass, we have analyzed the progression of replication on UV light-damaged DNA in mouse embryonic fibroblasts that contain a defined deletion in the N-terminal BRCT domain of Rev1 or that are deficient for Rev1. We provide evidence that Rev1 plays a coordinating role in two modes of DNA damage bypass, i.e., an early and a late pathway. The cells carrying the deletion in the BRCT domain are deficient for the early pathway, reflecting a role of the BRCT domain of Rev1 in mutagenic translesion synthesis. Rev1-deficient cells display a defect in both modes of DNA damage bypass. Despite the persistent defect in the late replicational bypass of fork-blocking (6-4)pyrimidine-pyrimidone photoproducts, overall replication is not strongly affected by Rev1 deficiency. This results in almost completely replicated templates that contain gaps encompassing the photoproducts. These gaps are inducers of DNA damage signaling leading to an irreversible G(2) arrest. Our results corroborate a model in which Rev1-mediated DNA damage bypass at postreplicative gaps quenches irreversible DNA damage responses.
Figures

Similar articles
-
The BRCT domain of mammalian Rev1 is involved in regulating DNA translesion synthesis.Nucleic Acids Res. 2005 Jan 13;33(1):356-65. doi: 10.1093/nar/gki189. Print 2005. Nucleic Acids Res. 2005. PMID: 15653636 Free PMC article.
-
Temporally distinct translesion synthesis pathways for ultraviolet light-induced photoproducts in the mammalian genome.DNA Repair (Amst). 2012 Jun 1;11(6):550-8. doi: 10.1016/j.dnarep.2012.03.007. Epub 2012 Apr 20. DNA Repair (Amst). 2012. PMID: 22521143
-
Translesion synthesis mechanisms depend on the nature of DNA damage in UV-irradiated human cells.Nucleic Acids Res. 2016 Jul 8;44(12):5717-31. doi: 10.1093/nar/gkw280. Epub 2016 Apr 19. Nucleic Acids Res. 2016. PMID: 27095204 Free PMC article.
-
Roles of mutagenic translesion synthesis in mammalian genome stability, health and disease.DNA Repair (Amst). 2015 May;29:56-64. doi: 10.1016/j.dnarep.2015.01.001. Epub 2015 Jan 21. DNA Repair (Amst). 2015. PMID: 25655219 Review.
-
The human intra-S checkpoint response to UVC-induced DNA damage.Carcinogenesis. 2010 May;31(5):751-65. doi: 10.1093/carcin/bgp230. Epub 2009 Sep 30. Carcinogenesis. 2010. PMID: 19793801 Free PMC article. Review.
Cited by
-
PCNA ubiquitination is important, but not essential for translesion DNA synthesis in mammalian cells.PLoS Genet. 2011 Sep;7(9):e1002262. doi: 10.1371/journal.pgen.1002262. Epub 2011 Sep 8. PLoS Genet. 2011. PMID: 21931560 Free PMC article.
-
Replication Protein A Prohibits Diffusion of the PCNA Sliding Clamp along Single-Stranded DNA.Biochemistry. 2017 Apr 4;56(13):1824-1835. doi: 10.1021/acs.biochem.6b01213. Epub 2017 Feb 28. Biochemistry. 2017. PMID: 28177605 Free PMC article.
-
Eukaryotic Translesion DNA Synthesis on the Leading and Lagging Strands: Unique Detours around the Same Obstacle.Chem Rev. 2017 Jun 28;117(12):7857-7877. doi: 10.1021/acs.chemrev.7b00046. Epub 2017 May 12. Chem Rev. 2017. PMID: 28497687 Free PMC article. Review.
-
Temozolomide resistance mechanisms: unveiling the role of translesion DNA polymerase kappa in glioblastoma spheroids in vitro.Biosci Rep. 2024 May 29;44(5):BSR20230667. doi: 10.1042/BSR20230667. Biosci Rep. 2024. PMID: 38717250 Free PMC article.
-
Lead compound profiling for small molecule inhibitors of the REV1-CT/RIR Translesion synthesis Protein-Protein interaction.Bioorg Med Chem. 2024 May 15;106:117755. doi: 10.1016/j.bmc.2024.117755. Epub 2024 May 9. Bioorg Med Chem. 2024. PMID: 38749343 Free PMC article.
References
-
- Akagi, J. I., C. Masutani, Y. Kataoka, T. Kan, E. Ohashi, T. Mori, H. Ohmori, and F. Hanaoka. 2009. Interaction with DNA polymerase eta is required for nuclear accumulation of REV1 and suppression of spontaneous mutations in human cells. DNA Repair 8585-599. - PubMed
-
- Andersen, P. L., F. Xu, and W. Xiao. 2008. Eukaryotic DNA damage tolerance and translesion synthesis through covalent modifications of PCNA. Cell Res. 18162-173. - PubMed
-
- Avkin, S., M. Goldsmith, S. Velasco-Miguel, N. E. Geacintov, E. C. Friedberg, and Z. Livneh. 2004. Quantitative analysis of translesion DNA synthesis across a benzo[a]pyrene-guanine adduct in mammalian cells: the role of DNA polymerase kappa. J. Biol. Chem. 27953298-53305. - PubMed
-
- Avkin, S., Z. Sevilya, L. Toube, N. E. Geacintov, S. G. Chaney, M. Oren, and Z. Livneh. 2006. p53 and p21 regulate error-prone DNA repair to yield a lower mutation load. Mol. Cell 22407-413. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Molecular Biology Databases
