Cooperation and virulence of clinical Pseudomonas aeruginosa populations
- PMID: 19332772
- PMCID: PMC2669332
- DOI: 10.1073/pnas.0811741106
Cooperation and virulence of clinical Pseudomonas aeruginosa populations
Abstract
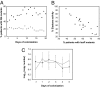
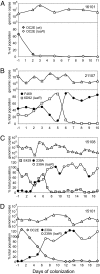
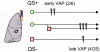
Bacteria communicate and cooperate to perform a wide range of social behaviors including production of extracellular products (public goods) that are crucial for growth and virulence. Their expression may be switched on by the detection of threshold densities of diffusible signals [Quorum-Sensing (QS)]. Studies using the opportunistic pathogen Pseudomonas aeruginosa suggest that QS "cheats"-individuals that don't respond to the QS signal, but are still able to use public goods produced by others-have a selective advantage in the presence of QS cooperators. It is, however, unclear whether this type of social exploitation is relevant in clinical contexts. Here, we report the evolutionary dynamics and virulence of P. aeruginosa populations during lung colonization of mechanically ventilated patients in the absence of antimicrobial treatments. We observed a large diversity of QS phenotypes among initial colonizing isolates. This diversity decreased over a matter of days, concomitant with a gradual increase in the proportion of QS cheating mutants (lasR mutants), which were found in 80% of the patients after 9 days of colonization. These mutants often evolved from initial wild-type genotypes. The fitness advantage of the lasR mutants is almost certainly due to social exploitation, because this advantage was only apparent in the presence of QS wild-type cells. Crucially, ventilator-associated pneumonia occurred significantly earlier in patients predominantly colonized by QS wild-type populations, highlighting the importance of QS in this clinical situation. These results demonstrate that social interactions can shape the short-term evolution and virulence of bacterial pathogens in humans, providing novel opportunities for therapy.
Conflict of interest statement
The authors declare no conflict of interest.
Figures

References
Publication types
MeSH terms
LinkOut - more resources
Full Text Sources

