Generation of transgene-free induced pluripotent mouse stem cells by the piggyBac transposon
- PMID: 19337237
- PMCID: PMC2677165
- DOI: 10.1038/nmeth.1323
Generation of transgene-free induced pluripotent mouse stem cells by the piggyBac transposon
Abstract
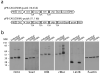
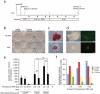
Induced pluripotent stem cells (iPSCs) have been generated from somatic cells by transgenic expression of Oct4 (Pou5f1), Sox2, Klf4 and Myc. A major difficulty in the application of this technology for regenerative medicine, however, is the delivery of reprogramming factors. Whereas retroviral transduction increases the risk of tumorigenicity, transient expression methods have considerably lower reprogramming efficiencies. Here we describe an efficient piggyBac transposon-based approach to generate integration-free iPSCs. Transposons carrying 2A peptide-linked reprogramming factors induced reprogramming of mouse embryonic fibroblasts with equivalent efficiencies to retroviral transduction. We removed transposons from these primary iPSCs by re-expressing transposase. Transgene-free iPSCs could be identified by negative selection. piggyBac excised without a footprint, leaving the iPSC genome without any genetic alteration. iPSCs fulfilled all criteria of pluripotency, such as pluripotency gene expression, teratoma formation and contribution to chimeras. piggyBac transposon-based reprogramming may be used to generate therapeutically applicable iPSCs.
Figures



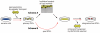
Comment in
-
Without a trace? PiggyBac-ing toward pluripotency.Nat Methods. 2009 May;6(5):329-30. doi: 10.1038/nmeth0509-329. Nat Methods. 2009. PMID: 19404251 No abstract available.
References
-
- Yamanaka S. Strategies and new developments in the generation of patient-specific pluripotent stem cells. Cell Stem Cell. 2007;1:39–49. - PubMed
-
- Takahashi K, Yamanaka S. Induction of pluripotent stem cells from mouse embryonic and adult fibroblast cultures by defined factors. Cell. 2006;126:663–676. - PubMed
-
- Takahashi K, et al. Induction of pluripotent stem cells from adult human fibroblasts by defined factors. Cell. 2007;131:861–872. - PubMed
-
- Yu J, et al. Induced pluripotent stem cell lines derived from human somatic cells. Science. 2007;318:1917–1920. - PubMed
-
- Okita K, Ichisaka T, Yamanaka S. Generation of germline-competent induced pluripotent stem cells. Nature. 2007;448:313–317. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources

