An agglomerative hierarchical approach to visualization in Bayesian clustering problems
- PMID: 19337306
- PMCID: PMC2705916
- DOI: 10.1038/hdy.2009.29
An agglomerative hierarchical approach to visualization in Bayesian clustering problems
Abstract
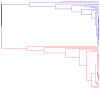
Clustering problems (including the clustering of individuals into outcrossing populations, hybrid generations, full-sib families and selfing lines) have recently received much attention in population genetics. In these clustering problems, the parameter of interest is a partition of the set of sampled individuals--the sample partition. In a fully Bayesian approach to clustering problems of this type, our knowledge about the sample partition is represented by a probability distribution on the space of possible sample partitions. As the number of possible partitions grows very rapidly with the sample size, we cannot visualize this probability distribution in its entirety, unless the sample is very small. As a solution to this visualization problem, we recommend using an agglomerative hierarchical clustering algorithm, which we call the exact linkage algorithm. This algorithm is a special case of the maximin clustering algorithm that we introduced previously. The exact linkage algorithm is now implemented in our software package PartitionView. The exact linkage algorithm takes the posterior co-assignment probabilities as input and yields as output a rooted binary tree, or more generally, a forest of such trees. Each node of this forest defines a set of individuals, and the node height is the posterior co-assignment probability of this set. This provides a useful visual representation of the uncertainty associated with the assignment of individuals to categories. It is also a useful starting point for a more detailed exploration of the posterior distribution in terms of the co-assignment probabilities.
Figures






Similar articles
-
A Bayesian approach to the identification of panmictic populations and the assignment of individuals.Genet Res. 2001 Aug;78(1):59-77. doi: 10.1017/s001667230100502x. Genet Res. 2001. PMID: 11556138
-
Variational Bayesian phylogenies through matrix representation of tree space.PeerJ. 2024 Apr 29;12:e17276. doi: 10.7717/peerj.17276. eCollection 2024. PeerJ. 2024. PMID: 38699195 Free PMC article.
-
Improving the inference of population genetic structure in the presence of related individuals.Genet Res (Camb). 2014;96:e003. doi: 10.1017/S0016672314000068. Genet Res (Camb). 2014. PMID: 25022872 Free PMC article.
-
H-CLAP: hierarchical clustering within a linear array with an application in genetics.Stat Appl Genet Mol Biol. 2015 Apr;14(2):125-41. doi: 10.1515/sagmb-2013-0076. Stat Appl Genet Mol Biol. 2015. PMID: 25803088
-
Folic acid supplementation and malaria susceptibility and severity among people taking antifolate antimalarial drugs in endemic areas.Cochrane Database Syst Rev. 2022 Feb 1;2(2022):CD014217. doi: 10.1002/14651858.CD014217. Cochrane Database Syst Rev. 2022. PMID: 36321557 Free PMC article.
Cited by
-
A spatial dirichlet process mixture model for clustering population genetics data.Biometrics. 2011 Jun;67(2):381-90. doi: 10.1111/j.1541-0420.2010.01484.x. Epub 2010 Sep 3. Biometrics. 2011. PMID: 20825394 Free PMC article.
References
-
- Aigner M. Combinatorial theory. Springer-Verlag; New York: 1979.
-
- Almudevar A, Field C. Inference of single generation sibling relationships based on DNA markers. Journal of Agricultural Biology and Environmental statistics. 1999;4:136–165.
-
- Berger JO. Statistical Decision Theory and Bayesian Analysis. 2nd edition Springer-Verlag; New York: 1985.
-
- Celeux G, Hurn M, Robert CP. Computational and inferential difficulties with mixture posterior distributions. Journal of the American Statistical Association. 2000;95:957–970.
Publication types
MeSH terms
Grants and funding
- 206/D16977/BB_/Biotechnology and Biological Sciences Research Council/United Kingdom
- BBS/E/C/00004937/BB_/Biotechnology and Biological Sciences Research Council/United Kingdom
- BBS/E/C/00004940/BB_/Biotechnology and Biological Sciences Research Council/United Kingdom
- D16977/BB_/Biotechnology and Biological Sciences Research Council/United Kingdom
- BBS/E/C/00004401/BB_/Biotechnology and Biological Sciences Research Council/United Kingdom
LinkOut - more resources
Full Text Sources
Research Materials
Miscellaneous

