ChIP-Seq of ERalpha and RNA polymerase II defines genes differentially responding to ligands
- PMID: 19339991
- PMCID: PMC2688537
- DOI: 10.1038/emboj.2009.88
ChIP-Seq of ERalpha and RNA polymerase II defines genes differentially responding to ligands
Abstract
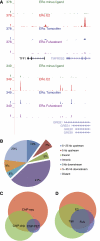
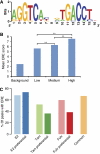
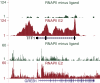
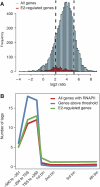
We used ChIP-Seq to map ERalpha-binding sites and to profile changes in RNA polymerase II (RNAPII) occupancy in MCF-7 cells in response to estradiol (E2), tamoxifen or fulvestrant. We identify 10 205 high confidence ERalpha-binding sites in response to E2 of which 68% contain an estrogen response element (ERE) and only 7% contain a FOXA1 motif. Remarkably, 596 genes change significantly in RNAPII occupancy (59% up and 41% down) already after 1 h of E2 exposure. Although promoter proximal enrichment of RNAPII (PPEP) occurs frequently in MCF-7 cells (17%), it is only observed on a minority of E2-regulated genes (4%). Tamoxifen and fulvestrant partially reduce ERalpha DNA binding and prevent RNAPII loading on the promoter and coding body on E2-upregulated genes. Both ligands act differently on E2-downregulated genes: tamoxifen acts as an agonist thus downregulating these genes, whereas fulvestrant antagonizes E2-induced repression and often increases RNAPII occupancy. Furthermore, our data identify genes preferentially regulated by tamoxifen but not by E2 or fulvestrant. Thus (partial) antagonist loaded ERalpha acts mechanistically different on E2-activated and E2-repressed genes.
Figures
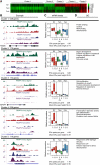
References
-
- Ali S, Coombes RC (2002) Endocrine-responsive breast cancer and strategies for combating resistance. Nat Rev Cancer 2: 101–112 - PubMed
-
- Cannings E, Kirkegaard T, Tovey SM, Dunne B, Cooke TG, Bartlett JM (2007) Bad expression predicts outcome in patients treated with tamoxifen. Breast Cancer Res Treat 102: 173–179 - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases

