Applications of metabolomics and proteomics to the mdx mouse model of Duchenne muscular dystrophy: lessons from downstream of the transcriptome
- PMID: 19341503
- PMCID: PMC2664943
- DOI: 10.1186/gm32
Applications of metabolomics and proteomics to the mdx mouse model of Duchenne muscular dystrophy: lessons from downstream of the transcriptome
Abstract
Functional genomic studies are dominated by transcriptomic approaches, in part reflecting the vast amount of information that can be obtained, the ability to amplify mRNA and the availability of commercially standardized functional genomic DNA microarrays and related techniques. This can be contrasted with proteomics, metabolomics and metabolic flux analysis (fluxomics), which have all been much slower in development, despite these techniques each providing a unique viewpoint of what is happening in the overall biological system. Here, we give an overview of developments in these fields 'downstream' of the transcriptome by considering the characterization of one particular, but widely used, mouse model of human disease. The mdx mouse is a model of Duchenne muscular dystrophy (DMD) and has been widely used to understand the progressive skeletal muscle wasting that accompanies DMD, and more recently the associated cardiomyopathy, as well as to unravel the roles of the other isoforms of dystrophin, such as those found in the brain. Studies using proteomics, metabolomics and fluxomics have characterized perturbations in calcium homeostasis in dystrophic skeletal muscle, provided an understanding of the role of dystrophin in skeletal muscle regeneration, and defined the changes in substrate energy metabolism in the working heart. More importantly, they all point to perturbations in proteins, metabolites and metabolic fluxes reflecting mitochondrial energetic alterations, even in the early stage of the dystrophic pathology. Philosophically, these studies also illustrate an important lesson relevant to both functional genomics and the mouse phenotyping in that the knowledge generated has advanced our understanding of cell biology and physiological organization as much as it has advanced our understanding of the disease.
Figures

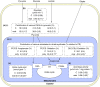
References
-
- Waterston RH, Lindblad-Toh K, Birney E, Rogers J, Abril JF, Agarwal P, Agarwala R, Ainscough R, Alexandersson M, An P, Antonarakis SE, Attwood J, Baertsch R, Bailey J, Barlow K, Beck S, Berry E, Birren B, Bloom T, Bork P, Botcherby M, Bray N, Brent MR, Brown DG, Brown SD, Bult C, Burton J, Butler J, Campbell RD, Carninci P, Cawley S. Initial sequencing and comparative analysis of the mouse genome. Nature. 2002;420:520–562. - PubMed
-
- Lander ES, Linton LM, Birren B, Nusbaum C, Zody MC, Baldwin J, Devon K, Dewar K, Doyle M, FitzHugh W, Funke R, Gage D, Harris K, Heaford A, Howland J, Kann L, Lehoczky J, LeVine R, McEwan P, McKernan K, Meldrim J, Mesirov JP, Miranda C, Morris W, Naylor J, Raymond C, Rosetti M, Santos R, Sheridan A, Sougnez C. Initial sequencing and analysis of the human genome. Nature. 2001;409:860–921. - PubMed
-
- Oliver SG, Winson MK, Kell DB, Baganz F. Systematic functional analysis of the yeast genome. Trends Biotech. 1998;16:373–378. - PubMed
-
- Raamsdonk LM, Teusink B, Broadhurst D, Zhang N, Hayes A, Walsh MC, Berden JA, Brindle KM, Kell DB, Rowland JJ, Westerhoff HV, van Dam K, Oliver SG. A functional genomics strategy that uses metabolome data to reveal the phenotype of silent mutations. Nat Biotechnol. 2001;19:45–50. - PubMed
-
- Castrillo JI, Hayes A, Mohammed S, Gaskell SJ, Oliver SG. An optimized protocol for metabolome analysis in yeast using direct infusion electrospray mass spectrometry. Phytochemistry. 2003;62:929–937. - PubMed
LinkOut - more resources
Full Text Sources

