Multiple-locus variable number tandem repeat analysis of Staphylococcus aureus: comparison with pulsed-field gel electrophoresis and spa-typing
- PMID: 19343175
- PMCID: PMC2661140
- DOI: 10.1371/journal.pone.0005082
Multiple-locus variable number tandem repeat analysis of Staphylococcus aureus: comparison with pulsed-field gel electrophoresis and spa-typing
Abstract
Background: Molecular typing of methicillin-resistant Staphylococcus aureus (MRSA) is required to study the routes and rates of transmission of this pathogen. Currently available typing techniques are either resource-intensive or have limited discriminatory ability. Multiple-locus variable number tandem repeat analysis (MLVA) may provide an alternative high throughput molecular typing tool with high epidemiological resolution.
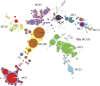
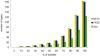
Methodology/principal findings: A new MLVA scheme for S. aureus was validated using 1681 S. aureus isolates collected from Dutch patients and 100 isolates from pigs. MLVA using 8 tandem repeat loci was performed in 2 multiplex PCRs and the fluorescently labeled PCR products were accurately sized on an automated DNA sequencer. The assessed number of repeats was used to create MLVA profiles consisting of strings of 8 integers that were used for categorical clustering. MLVA yielded 511 types that clustered into 11 distinct MLVA complexes which appeared to coincide with MLST clonal complexes. MLVA was at least as discriminatory as PFGE and twice as discriminatory as spa-sequence typing. There was considerable congruence between MLVA, spa-sequence typing and PFGE, at the MLVA complex level with group separation values of 95.1% and 89.2%. MLVA could not discriminate between pig-related MRSA strains isolated from humans and pigs, corroborating the high degree of relationship. MLVA was also superior in the grouping of MRSA isolates previously assigned to temporal-spatial clusters with indistinguishable SpaTypes, demonstrating its enhanced epidemiological usefulness.
Conclusions: The MLVA described in this study is a high throughput, relatively low cost genotyping method for S. aureus that yields discrete and unambiguous data that can be used to assign biological meaningful genotypes and complexes and can be used for interlaboratory comparisons in network accessible databases. Results suggest that MLVA offsets the disadvantages of other high discriminatory typing approaches and represents a promising tool for hospital, national and international molecular epidemiology.
Conflict of interest statement
Figures


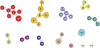

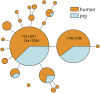
References
-
- Steinberg JP, Clark CC, Hackman BO. Nosocomial and community-acquired Staphylococcus aureus bacteremias from 1980 to 1993: impact of intravascular devices and methicillin resistance. Clin Infect Dis. 1996;23:255–259. - PubMed
-
- Panlilio AL, Culver DH, Gaynes RP, Banerjee S, Henderson TS, et al. Methicillin-resistant Staphylococcus aureus in U.S. hospitals, 1975–1991. Infect Control Hosp Epidemiol. 1992;13:582–586. - PubMed
-
- Speller DC, Johnson AP, James D, Marples RR, Charlett A, et al. Resistance to methicillin and other antibiotics in isolates of Staphylococcus aureus from blood and cerebrospinal fluid, England and Wales, 1989–95. Lancet. 1997;350:323–325. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Medical

