Quantitative proteomics reveals the function of unconventional ubiquitin chains in proteasomal degradation
- PMID: 19345192
- PMCID: PMC2668214
- DOI: 10.1016/j.cell.2009.01.041
Quantitative proteomics reveals the function of unconventional ubiquitin chains in proteasomal degradation
Abstract
All seven lysine residues in ubiquitin contribute to the synthesis of polyubiquitin chains on protein substrates. Whereas K48-linked chains are well established as mediators of proteasomal degradation, and K63-linked chains act in nonproteolytic events, the roles of unconventional polyubiquitin chains linked through K6, K11, K27, K29, or K33 are not well understood. Here, we report that the unconventional linkages are abundant in vivo and that all non-K63 linkages may target proteins for degradation. Ubiquitin with K48 as the single lysine cannot support yeast viability, and different linkages have partially redundant functions. By profiling both the entire yeast proteome and ubiquitinated proteins in wild-type and ubiquitin K11R mutant strains using mass spectrometry, we identified K11 linkage-specific substrates, including Ubc6, a ubiquitin-conjugating enzyme involved in endoplasmic reticulum-associated degradation (ERAD). Ubc6 primarily synthesizes K11-linked chains, and K11 linkages function in the ERAD pathway. Thus, unconventional polyubiquitin chains are critical for ubiquitin-proteasome system function.
Figures


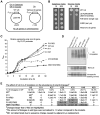




References
-
- Amerik AY, Li SJ, Hochstrasser M. Analysis of the deubiquitinating enzymes of the yeast Saccharomyces cerevisiae. Biol Chem. 2000a;381:981–992. - PubMed
-
- Baboshina OV, Haas AL. Novel multiubiquitin chain linkages catalyzed by the conjugating enzymes E2EPF and RAD6 are recognized by 26 S proteasome subunit 5. J Biol Chem. 1996;271:2823–2831. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases
Research Materials

