Cymbidium mosaic potexvirus isolate-dependent host movement systems reveal two movement control determinants and the coat protein is the dominant
- PMID: 19345971
- PMCID: PMC7103407
- DOI: 10.1016/j.virol.2009.02.049
Cymbidium mosaic potexvirus isolate-dependent host movement systems reveal two movement control determinants and the coat protein is the dominant
Abstract
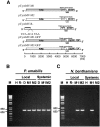
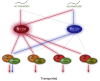
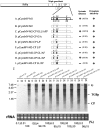
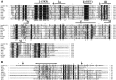
Little is known about how plant viruses of a single species exhibit different movement behavior in different host species. Two Cymbidium mosaic potexvirus (CymMV) isolates, M1 and M2, were studied. Both can infect Phalaenopsis orchids, but only M1 can systemically infect Nicotiana benthamiana plants. Protoplast inoculation and whole-mount in situ hybridization revealed that both isolates can replicate in N. benthamiana; however, M2 was restricted to the initially infected cells. Genome shuffling between M1 and M2 revealed that two control modes are involved in CymMV host dependent movement. The M1 coat protein (CP) plays a dominant role in controlling CymMV movement between cells, because all chimeric CymMV viruses containing the M1 CP systemically infected N. benthamiana plants. Without the M1 CP, one chimeric virus containing the combination of the M1 triple gene block proteins (TGBps), the M2 5' RNA (1-4333), and the M2 CP effectively moved in N. benthamiana plants. Further complementation analysis revealed that M1 TGBp1 and TGBp3 are co-required to complement the movement of the chimeric viruses in N. benthamiana. The amino acids within the CP, TGBp1 and TGBp3 which are required or important for CymMV M2 movement in N. benthamiana plants were mapped. The required amino acids within the CP map to the predicted RNA binding domain. RNA-protein binding assays revealed that M1 CP has higher RNA binding affinity than does M2 CP. Yeast two-hybrid assays to detect all possible interactions of M1 TGBps and CP, and only TGBp1 and CP self-interactions were observed.
Figures





Similar articles
-
Exploring the Multifunctional Roles of Odontoglossum Ringspot Virus P126 in Facilitating Cymbidium Mosaic Virus Cell-to-Cell Movement during Mixed Infection.Viruses. 2021 Aug 6;13(8):1552. doi: 10.3390/v13081552. Viruses. 2021. PMID: 34452417 Free PMC article.
-
Reciprocal function of movement proteins and complementation of long-distance movement of Cymbidium mosaic virus RNA by Odontoglossum ringspot virus coat protein.J Gen Virol. 2005 May;86(Pt 5):1543-1553. doi: 10.1099/vir.0.80772-0. J Gen Virol. 2005. PMID: 15831968
-
Potato virus X TGBp1 induces plasmodesmata gating and moves between cells in several host species whereas CP moves only in N. benthamiana leaves.Virology. 2004 Oct 25;328(2):185-97. doi: 10.1016/j.virol.2004.06.039. Virology. 2004. PMID: 15464839
-
Viral elements and host cellular proteins in intercellular movement of Bamboo mosaic virus.Curr Opin Virol. 2015 Jun;12:99-108. doi: 10.1016/j.coviro.2015.04.005. Epub 2015 May 13. Curr Opin Virol. 2015. PMID: 25951346 Review.
-
Molecular biology of potexviruses: recent advances.J Gen Virol. 2007 Jun;88(Pt 6):1643-1655. doi: 10.1099/vir.0.82667-0. J Gen Virol. 2007. PMID: 17485523 Review.
Cited by
-
Two key arginine residues in the coat protein of Bamboo mosaic virus differentially affect the accumulation of viral genomic and subgenomic RNAs.Mol Plant Pathol. 2014 Feb;15(2):196-210. doi: 10.1111/mpp.12080. Epub 2013 Oct 28. Mol Plant Pathol. 2014. PMID: 24393453 Free PMC article.
-
Genome-wide analysis of small RNAs from Odontoglossum ringspot virus and Cymbidium mosaic virus synergistically infecting Phalaenopsis.Mol Plant Pathol. 2020 Feb;21(2):188-205. doi: 10.1111/mpp.12888. Epub 2019 Nov 14. Mol Plant Pathol. 2020. PMID: 31724809 Free PMC article.
-
The Coat Protein of Citrus Yellow Vein Clearing Virus Interacts with Viral Movement Proteins and Serves as an RNA Silencing Suppressor.Viruses. 2019 Apr 5;11(4):329. doi: 10.3390/v11040329. Viruses. 2019. PMID: 30959816 Free PMC article.
-
Exploring the Multifunctional Roles of Odontoglossum Ringspot Virus P126 in Facilitating Cymbidium Mosaic Virus Cell-to-Cell Movement during Mixed Infection.Viruses. 2021 Aug 6;13(8):1552. doi: 10.3390/v13081552. Viruses. 2021. PMID: 34452417 Free PMC article.
References
-
- Abouhaidar M.G., Lai R. Nucleotide sequence of the 3′-terminal region of clover yellow mosaic virus RNA. J. Gen. Virol. 1989;70:1871–1875. - PubMed
-
- Adams M.J., Accotto G.P., Agranovsky A.A., Bar-Joseph M., Boscia D., Brunt A.A., Candresse T., Coutts R.H.A., Dolja V.V., Falk B.W., Foster G.D., Gonsalves D., Jelkmann W., Karasev A., Martelli G.P., Mawassi M., Milne R.G., Minafra A., Namba S., Rowhani A., Vetten H.J., Vishnichenko V.K., Wisler G.C., Yoshikawa N., Zavriev S.K. Genus Potexvirus. In: Fauquet C.M., Mayo M.A., Maniloff J., Desselberger U., Ball L.A., editors. Virus Taxonomy: Eighth Report of the International Committee on Taxonomy of Virus. Elsevier Academic Press; San Diego, CA: 2005. pp. 1091–1095.
-
- Adams M.J., Accotto G.P., Agranovsky A.A., Bar-Joseph M., Boscia D., Brunt A.A., Candresse T., Coutts R.H.A., Dolja V.V., Falk B.W., Foster G.D., Gonsalves D., Jelkmann W., Karasev A., Martelli G.P., Mawassi M., Milne R.G., Minafra A., Namba S., Rowhani A., Vetten H.J., Vishnichenko V.K., Wisler G.C., Yoshikawa N., Zavriev S.K. Flexiviridae. In: Fauquet C.M., Mayo M.A., Maniloff J., Desselberger U., Ball L.A., editors. Fauquet, Virus Taxonomy : Eight Report of the International Committee on Taxonomy of Viruses. Elsevier Academic Press; San Diego, CA: 2005. pp. 1101–1124.
-
- Angell S.M., Davies C., Baulcombe D.C. Cell-to-cell movement of potato virus X is associated with a change in the size-exclusion limit of plasmodesmata in trichome cells of Nicotiana clevelandii. Virology. 1996;216:197–201. - PubMed
-
- Bayne E.H., Rakitina D.V., Morozov S.Y.u., Baulcombe D.C. Cell-to-cell movement of potato potexvirus X is dependent on suppression of RNA silencing. Plant J. 2005;44:471–482. - PubMed
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Miscellaneous

