Characterization of a unique class C acid phosphatase from Clostridium perfringens
- PMID: 19363079
- PMCID: PMC2687270
- DOI: 10.1128/AEM.01599-08
Characterization of a unique class C acid phosphatase from Clostridium perfringens
Abstract
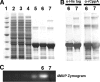
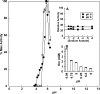
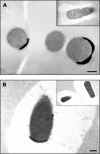
Clostridium perfringens is a gram-positive anaerobe and a pathogen of medical importance. The detection of acid phosphatase activity is a powerful diagnostic indicator of the presence of C. perfringens among anaerobic isolates; however, characterization of the enzyme has not previously been reported. Provided here are details of the characterization of a soluble recombinant form of this cell-associated enzyme. The denatured enzyme was approximately 31 kDa and a homodimer in solution. It catalyzed the hydrolysis of several substrates, including para-nitrophenyl phosphate, 4-methylumbelliferyl phosphate, and 3' and 5' nucleoside monophosphates at pH 6. Calculated K(m)s ranged from 0.2 to 0.6 mM with maximum velocity ranging from 0.8 to 1.6 micromol of P(i)/s/mg. Activity was enhanced in the presence of some divalent cations but diminished in the presence of others. Wild-type enzyme was detected in all clinical C. perfringens isolates tested and found to be cell associated. The described enzyme belongs to nonspecific acid phosphatase class C but is devoid of lipid modification commonly attributed to this class.
Figures

References
-
- Bateman, A., and M. Bycroft. 2000. The structure of a LysM domain from E. coli membrane-bound lytic murein transglycosylase D (MltD). J. Mol. Biol. 299:1113-1119. - PubMed
-
- Calderone, V., C. Forleo, M. Benvenuti, M. C. Thaller, G. M. Rossolini, and S. Mangani. 2006. The first structure of a bacterial class B acid phosphatase reveals further structural heterogeneity among phosphatases of the haloacid dehalogenase fold. J. Mol. Biol. 365:708-721. - PubMed
-
- Chance, D. L., C. A. Jensen, T. J. Reilly, and T. P. Mawhinney. 2005. Analysis of bacterial colony biology using microwave-assisted processing and LR white resin. Microsc. Microanal. 11(Suppl. 2):968-969.
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Molecular Biology Databases

