Advanced backcross-QTL analysis in spring barley (H. vulgare ssp. spontaneum) comparing a REML versus a Bayesian model in multi-environmental field trials
- PMID: 19363603
- PMCID: PMC2755740
- DOI: 10.1007/s00122-009-1021-6
Advanced backcross-QTL analysis in spring barley (H. vulgare ssp. spontaneum) comparing a REML versus a Bayesian model in multi-environmental field trials
Abstract
A common difficulty in mapping quantitative trait loci (QTLs) is that QTL effects may show environment specificity and thus differ across environments. Furthermore, quantitative traits are likely to be influenced by multiple QTLs or genes having different effect sizes. There is currently a need for efficient mapping strategies to account for both multiple QTLs and marker-by-environment interactions. Thus, the objective of our study was to develop a Bayesian multi-locus multi-environmental method of QTL analysis. This strategy is compared to (1) Bayesian multi-locus mapping, where each environment is analysed separately, (2) Restricted Maximum Likelihood (REML) single-locus method using a mixed hierarchical model, and (3) REML forward selection applying a mixed hierarchical model. For this study, we used data on multi-environmental field trials of 301 BC(2)DH lines derived from a cross between the spring barley elite cultivar Scarlett and the wild donor ISR42-8 from Israel. The lines were genotyped by 98 SSR markers and measured for the agronomic traits "ears per m(2)," "days until heading," "plant height," "thousand grain weight," and "grain yield". Additionally, a simulation study was performed to verify the QTL results obtained in the spring barley population. In general, the results of Bayesian QTL mapping are in accordance with REML methods. In this study, Bayesian multi-locus multi-environmental analysis is a valuable method that is particularly suitable if lines are cultivated in multi-environmental field trials.
Figures

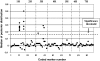
References
-
- Bauer AM, Reetz TC, Léon J (2006) Estimation of breeding values of inbred lines using best linear unbiased prediction (BLUP) and genetic similarities. Crop Sci 46:2685–2691 - DOI
-
- Bauer AM, Reetz TC, Léon J (2008) Predicting breeding values of spring barley accessions by using the singular value decomposition of genetic similarities. Plant Breed 127:274–278 - DOI
-
- Bezant J, Laurie D, Pratchett N, Chojecki J, Kearsey M (1996) Marker regression mapping of QTL controlling flowering time and plant height in a spring barley (Hordeum vulgare L.) cross. Heredity 77:64–73 - DOI
-
- Boer MP, Wright D, Feng L, Podlich DW, Luo L, Cooper M, van Eeuwijk FA (2007) A mixed-model quantitative trait loci (QTL) analysis for multiple-environment trial data using environmental covariables for QTL-by-environment interactions, with an example in maize. Genetics 177:1801–1813 - DOI - PMC - PubMed
Publication types
MeSH terms
LinkOut - more resources
Full Text Sources

