Conservation of core gene expression in vertebrate tissues
- PMID: 19371447
- PMCID: PMC2689434
- DOI: 10.1186/jbiol130
Conservation of core gene expression in vertebrate tissues
Abstract
Background: Vertebrates share the same general body plan and organs, possess related sets of genes, and rely on similar physiological mechanisms, yet show great diversity in morphology, habitat and behavior. Alteration of gene regulation is thought to be a major mechanism in phenotypic variation and evolution, but relatively little is known about the broad patterns of conservation in gene expression in non-mammalian vertebrates.
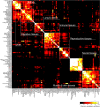
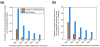
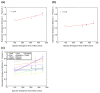
Results: We measured expression of all known and predicted genes across twenty tissues in chicken, frog and pufferfish. By combining the results with human and mouse data and considering only ten common tissues, we have found evidence of conserved expression for more than a third of unique orthologous genes. We find that, on average, transcription factor gene expression is neither more nor less conserved than that of other genes. Strikingly, conservation of expression correlates poorly with the amount of conserved nonexonic sequence, even using a sequence alignment technique that accounts for non-collinearity in conserved elements. Many genes show conserved human/fish expression despite having almost no nonexonic conserved primary sequence.
Conclusions: There are clearly strong evolutionary constraints on tissue-specific gene expression. A major challenge will be to understand the precise mechanisms by which many gene expression patterns remain similar despite extensive cis-regulatory restructuring.
Figures





References
-
- Jaillon O, Aury JM, Brunet F, Petit JL, Stange-Thomann N, Mauceli E, Bouneau L, Fischer C, Ozouf-Costaz C, Bernot A, Nicaud S, Jaffe D, Fisher S, Lutfalla G, Dossat C, Segurens B, Dasilva C, Salanoubat M, Levy M, Boudet N, Castellano S, Anthouard V, Jubin C, Castelli V, Katinka M, Vacherie B, Biémont C, Skalli Z, Cattolico L, Poulain J, et al. Genome duplication in the teleost fish Tetraodon nigroviridis reveals the early vertebrate protokaryotype. Nature. 2004;431:946–957. doi: 10.1038/nature03025. - DOI - PubMed
-
- Mouse Genome Sequencing Consortium. Waterston RH, Lindblad-Toh K, Birney E, Rogers J, Abril JF, Agarwal P, Agarwala R, Ainscough R, Alexandersson M, An P, Antonarakis SE, Attwood J, Baertsch R, Bailey J, Barlow K, Beck S, Berry E, Birren B, Bloom T, Bork P, Botcherby M, Bray N, Brent MR, Brown DG, Brown SD, Bult C, Burton J, Butler J, Campbell RD, et al. Initial sequencing and comparative analysis of the mouse genome. Nature. 2002;420:520–562. doi: 10.1038/nature01262. - DOI - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases

