Requirements for E1A dependent transcription in the yeast Saccharomyces cerevisiae
- PMID: 19374760
- PMCID: PMC2674444
- DOI: 10.1186/1471-2199-10-32
Requirements for E1A dependent transcription in the yeast Saccharomyces cerevisiae
Abstract
Background: The human adenovirus type 5 early region 1A (E1A) gene encodes proteins that are potent regulators of transcription. E1A does not bind DNA directly, but is recruited to target promoters by the interaction with sequence specific DNA binding proteins. In mammalian systems, E1A has been shown to contain two regions that can independently induce transcription when fused to a heterologous DNA binding domain. When expressed in Saccharomyces cerevisiae, each of these regions of E1A also acts as a strong transcriptional activator. This allows yeast to be used as a model system to study mechanisms by which E1A stimulates transcription.
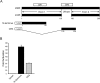
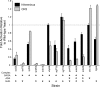
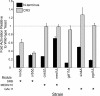
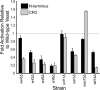
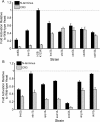
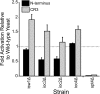
Results: Using 81 mutant yeast strains, we have evaluated the effect of deleting components of the ADA, COMPASS, CSR, INO80, ISW1, NuA3, NuA4, Mediator, PAF, RSC, SAGA, SAS, SLIK, SWI/SNF and SWR1 transcriptional regulatory complexes on E1A dependent transcription. In addition, we examined the role of histone H2B ubiquitylation by Rad6/Bre1 on transcriptional activation.
Conclusion: Our analysis indicates that the two activation domains of E1A function via distinct mechanisms, identify new factors regulating E1A dependent transcription and suggest that yeast can serve as a valid model system for at least some aspects of E1A function.
Figures
References
-
- Bayley ST, Mymryk JS. Adenovirus E1A proteins and transformation (Review) Int J Oncology. 1994;5:425–444. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Molecular Biology Databases
Research Materials

