How are T(H)1 and T(H)2 effector cells made?
- PMID: 19375293
- PMCID: PMC2695256
- DOI: 10.1016/j.coi.2009.03.010
How are T(H)1 and T(H)2 effector cells made?
Abstract
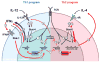
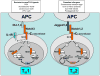
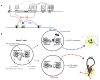
Differentiation of T(H)1 and T(H)2 effector cells proceeds through several phases: First, naïve CD4(+) precursor cells are instructed to differentiate as appropriate to optimally fight the infectious threat encountered. This process is governed by the IL12 and IL4 cytokines, as well as by signaling through the Notch receptor. In response to these signals, transcription is initiated of lineage specific cytokine genes including the Ifngamma and Il4 genes as well as of genes encoding transcriptional regulators, such as T-bet and Gata3. The respective differentiation programs are reinforced by both positive and negative feedback mechanisms. Furthermore, epigenetic modifications of the lineage specific genes result in the emergence of regulatory elements, which control high level lineage restricted expression by both intrachromosomal and interchromosomal associations. Together, these mechanisms ensure stable inheritance of the differentiated fate in the numerous progeny of the original naïve CD4(+) T cells.
Figures

References
-
- Abbas AK, Murphy KM, Sher A. Functional diversity of helper T lymphocytes. Nature. 1996;383:787–793. - PubMed
-
- Kapsenberg ML. Dendritic-cell control of pathogen-driven T-cell polarization. Nat Rev Immunol. 2003;3:984–993. - PubMed
-
- Thieu VT, Yu Q, Chang HC, Yeh N, Nguyen ET, Sehra S, Kaplan MH. Signal transducer and activator of transcription 4 is required for the transcription factor T-bet to promote T helper 1 cell-fate determination . Immunity. 2008;29:679–690. Rather than having a linear relationship, STAT4 and T-bet are shown to have complementary roles in regulation of the TH1 transcriptional program, with some genes dependent on one and other genes dependent on both transcription factors. - PMC - PubMed
-
- Weaver CT, Hatton RD, Mangan PR, Harrington LE. IL-17 family cytokines and the expanding diversity of effector T cell lineages. Annu Rev Immunol. 2007;25:821–852. - PubMed
-
- Djuretic IM, Levanon D, Negreanu V, Groner Y, Rao A, Ansel KM. Transcription factors T-bet and Runx3 cooperate to activate Ifng and silence Il4 in T helper type 1 cells. Nat Immunol. 2007;8:145–153. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Research Materials

