Different fecal microbiotas and volatile organic compounds in treated and untreated children with celiac disease
- PMID: 19376912
- PMCID: PMC2698361
- DOI: 10.1128/AEM.02793-08
Different fecal microbiotas and volatile organic compounds in treated and untreated children with celiac disease
Abstract
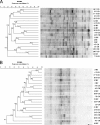
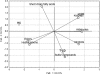
This study aimed at investigating the fecal microbiotas of children with celiac disease (CD) before (U-CD) and after (T-CD) they were fed a gluten-free diet and of healthy children (HC). Brothers or sisters of T-CD were enrolled as HC. Each group consisted of seven children. PCR-denaturing gradient gel electrophoresis (DGGE) analysis with V3 universal primers revealed a unique profile for each fecal sample. PCR-DGGE analysis with group- or genus-specific 16S rRNA gene primers showed that the Lactobacillus community of U-CD changed significantly, while the diversity of the Lactobacillus community of T-CD was quite comparable to that of HC. Compared to HC, the ratio of cultivable lactic acid bacteria and Bifidobacterium to Bacteroides and enterobacteria was lower in T-CD and even lower in U-CD. The percentages of strains identified as lactobacilli differed as follows: HC (ca. 38%) > T-CD (ca. 17%) > U-CD (ca. 10%). Lactobacillus brevis, Lactobacillus rossiae, and Lactobacillus pentosus were identified only in fecal samples from T-CD and HC. Lactobacillus fermentum, Lactobacillus delbrueckii subsp. bulgaricus, and Lactobacillus gasseri were identified only in several fecal samples from HC. Compared to HC, the composition of Bifidobacterium species of T-CD varied, and it varied even more for U-CD. Forty-seven volatile organic compounds (VOCs) belonging to different chemical classes were identified using gas-chromatography mass spectrometry-solid-phase microextraction analysis. The median concentrations varied markedly for HC, T-CD, and U-CD. Overall, the r(2) values for VOC data for brothers and sisters were equal to or lower than those for unrelated HC and T-CD. This study shows the effect of CD pathology on the fecal microbiotas of children.
Figures


Similar articles
-
Duodenal and faecal microbiota of celiac children: molecular, phenotype and metabolome characterization.BMC Microbiol. 2011 Oct 4;11:219. doi: 10.1186/1471-2180-11-219. BMC Microbiol. 2011. PMID: 21970810 Free PMC article.
-
Temporal stability analysis of the microbiota in human feces by denaturing gradient gel electrophoresis using universal and group-specific 16S rRNA gene primers.FEMS Microbiol Ecol. 2004 Jun 1;48(3):437-46. doi: 10.1016/j.femsec.2004.03.001. FEMS Microbiol Ecol. 2004. PMID: 19712312
-
Salivary and fecal microbiota and metabolome of celiac children under gluten-free diet.Int J Food Microbiol. 2016 Dec 19;239:125-132. doi: 10.1016/j.ijfoodmicro.2016.07.025. Epub 2016 Jul 19. Int J Food Microbiol. 2016. PMID: 27452636 Review.
-
Salivary microbiota and metabolome associated with celiac disease.Appl Environ Microbiol. 2014 Jun;80(11):3416-25. doi: 10.1128/AEM.00362-14. Epub 2014 Mar 21. Appl Environ Microbiol. 2014. PMID: 24657864 Free PMC article.
-
Analytical Tools for Disease Diagnosis in Animals via Fecal Volatilome.Crit Rev Anal Chem. 2022;52(5):917-932. doi: 10.1080/10408347.2020.1843130. Epub 2020 Nov 12. Crit Rev Anal Chem. 2022. PMID: 33180561 Review.
Cited by
-
A low-gluten diet induces changes in the intestinal microbiome of healthy Danish adults.Nat Commun. 2018 Nov 13;9(1):4630. doi: 10.1038/s41467-018-07019-x. Nat Commun. 2018. PMID: 30425247 Free PMC article. Clinical Trial.
-
Faecal Scent as a Novel Non-Invasive Biomarker to Discriminate between Coeliac Disease and Refractory Coeliac Disease: A Proof of Principle Study.Biosensors (Basel). 2019 May 27;9(2):69. doi: 10.3390/bios9020069. Biosensors (Basel). 2019. PMID: 31137798 Free PMC article.
-
Draft genome sequence of Lactobacillus rossiae DSM 15814(T).J Bacteriol. 2012 Oct;194(19):5460-1. doi: 10.1128/JB.01248-12. J Bacteriol. 2012. PMID: 22965087 Free PMC article.
-
Cloning, expression and characterization of a β-D-xylosidase from Lactobacillus rossiae DSM 15814(T).Microb Cell Fact. 2016 May 3;15:72. doi: 10.1186/s12934-016-0473-z. Microb Cell Fact. 2016. PMID: 27142164 Free PMC article.
-
The Contribution of the Intestinal Microbiota to the Celiac Disease Pathogenesis along with the Effectiveness of Probiotic Therapy.Microorganisms. 2023 Nov 23;11(12):2848. doi: 10.3390/microorganisms11122848. Microorganisms. 2023. PMID: 38137992 Free PMC article. Review.
References
-
- Adlerberth, I., M. Cerquetti, I. Poilane, A. Wold, and A. Collignon. 2000. Mechanisms of colonisation and colonisation resistance of the digestive tract. Microb. Ecol. Health Dis. 11:223-239.
-
- Collado, M. C., M. Calabuig, and Y. Sanz. 2007. Differences between the fecal microbiota of celiac infants and healthy controls. Curr. Issues Intest. Microbiol. 8:9-14. - PubMed
-
- De Angelis, M., C. G. Rizzello, A. Fasano, M. G. Clemente, C. De Simone, M. Silano, M. De Vincenzi, I. Losito, and M. Gobbetti. 2005. VSL#3 probiotic preparation has the capacity to hydrolyze gliadin polypeptides responsible for celiac sprue. Biochim. Biophys. Acta 1762:80-93. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Medical
Molecular Biology Databases

