PIF3 is a repressor of chloroplast development
- PMID: 19380736
- PMCID: PMC2678601
- DOI: 10.1073/pnas.0811684106
PIF3 is a repressor of chloroplast development
Abstract
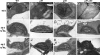
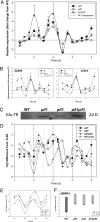
The phytochrome-interacting factor PIF3 has been proposed to act as a positive regulator of chloroplast development. Here, we show that the pif3 mutant has a phenotype that is similar to the pif1 mutant, lacking the repressor of chloroplast development PIF1, and that a pif1pif3 double mutant has an additive phenotype in all respects. The pif mutants showed elevated protochlorophyllide levels in the dark, and etioplasts of pif mutants contained smaller prolamellar bodies and more prothylakoid membranes than corresponding wild-type seedlings, similar to previous reports of constitutive photomorphogenic mutants. Consistent with this observation, pif1, pif3, and pif1pif3 showed reduced hypocotyl elongation and increased cotyledon opening in the dark. Transfer of 4-d-old dark-grown seedlings to white light resulted in more chlorophyll synthesis in pif mutants over the first 2 h, and analysis of gene expression in dark-grown pif mutants indicated that key tetrapyrrole regulatory genes such as HEMA1 encoding the rate-limiting step in tetrapyrrole synthesis were already elevated 2 d after germination. Circadian regulation of HEMA1 in the dark also showed reduced amplitude and a shorter, variable period in the pif mutants, whereas expression of the core clock components TOC1, CCA1, and LHY was largely unaffected. Expression of both PIF1 and PIF3 was circadian regulated in dark-grown seedlings. PIF1 and PIF3 are proposed to be negative regulators that function to integrate light and circadian control in the regulation of chloroplast development.
Conflict of interest statement
The authors declare no conflict of interest.
Figures


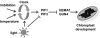
Similar articles
-
PIF1 promotes phytochrome-regulated growth under photoperiodic conditions in Arabidopsis together with PIF3, PIF4, and PIF5.J Exp Bot. 2014 Jun;65(11):2925-36. doi: 10.1093/jxb/ert465. Epub 2014 Jan 13. J Exp Bot. 2014. PMID: 24420574 Free PMC article.
-
Phytochrome-imposed oscillations in PIF3 protein abundance regulate hypocotyl growth under diurnal light/dark conditions in Arabidopsis.Plant J. 2012 Aug;71(3):390-401. doi: 10.1111/j.1365-313X.2012.04992.x. Epub 2012 Jun 11. Plant J. 2012. PMID: 22409654 Free PMC article.
-
Phytochromes promote seedling light responses by inhibiting four negatively-acting phytochrome-interacting factors.Proc Natl Acad Sci U S A. 2009 May 5;106(18):7660-5. doi: 10.1073/pnas.0812219106. Epub 2009 Apr 20. Proc Natl Acad Sci U S A. 2009. PMID: 19380720 Free PMC article.
-
Beyond the darkness: recent lessons from etiolation and de-etiolation studies.J Exp Bot. 2020 Feb 19;71(4):1215-1225. doi: 10.1093/jxb/erz496. J Exp Bot. 2020. PMID: 31854450 Free PMC article. Review.
-
Genetic dissection of chloroplast biogenesis and development: an overview.Plant Physiol. 2011 Apr;155(4):1545-51. doi: 10.1104/pp.110.170365. Epub 2011 Feb 17. Plant Physiol. 2011. PMID: 21330494 Free PMC article. Review. No abstract available.
Cited by
-
Antagonistic basic helix-loop-helix/bZIP transcription factors form transcriptional modules that integrate light and reactive oxygen species signaling in Arabidopsis.Plant Cell. 2013 May;25(5):1657-73. doi: 10.1105/tpc.112.104869. Epub 2013 May 3. Plant Cell. 2013. PMID: 23645630 Free PMC article.
-
Light signaling in plants-a selective history.Plant Physiol. 2024 Apr 30;195(1):213-231. doi: 10.1093/plphys/kiae110. Plant Physiol. 2024. PMID: 38431282 Free PMC article. Review.
-
Direct regulation of phytoene synthase gene expression and carotenoid biosynthesis by phytochrome-interacting factors.Proc Natl Acad Sci U S A. 2010 Jun 22;107(25):11626-31. doi: 10.1073/pnas.0914428107. Epub 2010 Jun 7. Proc Natl Acad Sci U S A. 2010. PMID: 20534526 Free PMC article.
-
EIN3/EIL1 cooperate with PIF1 to prevent photo-oxidation and to promote greening of Arabidopsis seedlings.Proc Natl Acad Sci U S A. 2009 Dec 15;106(50):21431-6. doi: 10.1073/pnas.0907670106. Epub 2009 Nov 30. Proc Natl Acad Sci U S A. 2009. PMID: 19948955 Free PMC article.
-
Phytochromes inhibit hypocotyl negative gravitropism by regulating the development of endodermal amyloplasts through phytochrome-interacting factors.Proc Natl Acad Sci U S A. 2011 Jan 25;108(4):1729-34. doi: 10.1073/pnas.1011066108. Epub 2011 Jan 10. Proc Natl Acad Sci U S A. 2011. PMID: 21220341 Free PMC article.
References
-
- Chen M, Chory J, Fankhauser C. Light signal transduction in higher plants. Annu Rev Genet. 2004;38:87–117. - PubMed
-
- Bae G, Choi G. Decoding of light signals by plant phytochromes and their interacting proteins. Annu Rev Plant Biol. 2008;59:281–311. - PubMed
-
- Duek PD, Fankhauser C. bHLH class transcription factors take centre stage in phytochrome signalling. Trends Plants Sci. 2005;10:51–54. - PubMed
-
- Ni M, Tepperman JM, Quail PH. PIF3, a phytochrome-interacting factor necessary for normal photoinduced signal transduction, is a novel basic helix-loop-helix protein. Cell. 1998;95:657–667. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Molecular Biology Databases

