Influence of whole-body dynamics on 15N PISEMA NMR spectra of membrane proteins: a theoretical analysis
- PMID: 19383467
- PMCID: PMC2718307
- DOI: 10.1016/j.bpj.2008.12.3950
Influence of whole-body dynamics on 15N PISEMA NMR spectra of membrane proteins: a theoretical analysis
Abstract
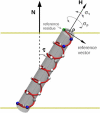
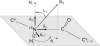
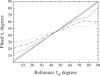
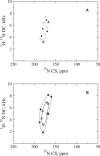
Membrane proteins and peptides exhibit a preferred orientation in the lipid bilayer while fluctuating in an anisotropic manner. Both the orientation and the dynamics have direct functional implications, but motions are usually not accessible, and structural descriptions are generally static. Using simulated data, we analyze systematically the impact of whole-body motions on the peptide orientations calculated from two-dimensional polarization inversion spin exchange at the magic angle (PISEMA) NMR. Fluctuations are found to have a significant effect on the observed spectra. Nevertheless, wheel-like patterns are still preserved, and it is possible to determine the average peptide tilt and azimuthal rotation angles using simple static models for the spectral fitting. For helical peptides undergoing large-amplitude fluctuations, as in the case of transmembrane monomers, improved fits can be achieved using an explicit dynamics model that includes Gaussian distributions of the orientational parameters. This method allows extracting the amplitudes of fluctuations of the tilt and azimuthal rotation angles. The analysis is further demonstrated by generating first a virtual PISEMA spectrum from a molecular dynamics trajectory of the model peptide, WLP23, in a lipid membrane. That way, the dynamics of the system from which the input spectrum originates is completely known at atomic detail and can thus be directly compared with the dynamic output obtained from the fit. We find that fitting our dynamics model to the polar index slant angles wheel gives an accurate description of the amplitude of underlying motions, together with the average peptide orientation.
Figures

Similar articles
-
Comparative analysis of the orientation of transmembrane peptides using solid-state (2)H- and (15)N-NMR: mobility matters.Eur Biophys J. 2012 May;41(5):475-82. doi: 10.1007/s00249-012-0801-0. Epub 2012 Mar 28. Eur Biophys J. 2012. PMID: 22453992
-
The dynamic orientation of membrane-bound peptides: bridging simulations and experiments.Biophys J. 2007 Dec 15;93(12):4278-88. doi: 10.1529/biophysj.107.113043. Epub 2007 Aug 24. Biophys J. 2007. PMID: 17720729 Free PMC article.
-
On the orientation of a designed transmembrane peptide: toward the right tilt angle?J Am Chem Soc. 2007 Dec 12;129(49):15174-81. doi: 10.1021/ja073784q. Epub 2007 Nov 15. J Am Chem Soc. 2007. PMID: 18001020
-
Structure determination of membrane proteins by NMR spectroscopy.Biochem Cell Biol. 2002;80(5):597-604. doi: 10.1139/o02-154. Biochem Cell Biol. 2002. PMID: 12440700 Free PMC article. Review.
-
Lipid bilayers: an essential environment for the understanding of membrane proteins.Magn Reson Chem. 2007 Dec;45 Suppl 1:S2-11. doi: 10.1002/mrc.2077. Epub 2007 Dec 19. Magn Reson Chem. 2007. PMID: 18095258 Review.
Cited by
-
Supramolecular structure of membrane-associated polypeptides by combining solid-state NMR and molecular dynamics simulations.Biophys J. 2012 Jul 3;103(1):29-37. doi: 10.1016/j.bpj.2012.05.016. Biophys J. 2012. PMID: 22828329 Free PMC article.
-
Comparative analysis of the orientation of transmembrane peptides using solid-state (2)H- and (15)N-NMR: mobility matters.Eur Biophys J. 2012 May;41(5):475-82. doi: 10.1007/s00249-012-0801-0. Epub 2012 Mar 28. Eur Biophys J. 2012. PMID: 22453992
-
Probing the transmembrane structure and dynamics of microsomal NADPH-cytochrome P450 oxidoreductase by solid-state NMR.Biophys J. 2014 May 20;106(10):2126-33. doi: 10.1016/j.bpj.2014.03.051. Biophys J. 2014. PMID: 24853741 Free PMC article.
-
Irregular structure of the HIV fusion peptide in membranes demonstrated by solid-state NMR and MD simulations.Eur Biophys J. 2011 Apr;40(4):529-43. doi: 10.1007/s00249-011-0676-5. Epub 2011 Jan 28. Eur Biophys J. 2011. PMID: 21274707
-
Comparison between the behavior of different hydrophobic peptides allowing membrane anchoring of proteins.Adv Colloid Interface Sci. 2014 May;207:223-39. doi: 10.1016/j.cis.2014.01.015. Epub 2014 Jan 28. Adv Colloid Interface Sci. 2014. PMID: 24560216 Free PMC article. Review.
References
-
- Singer S.J., Nicolson G.L. The fluid mosaic model of the structure of cell membranes. Science. 1972;175:720–731. - PubMed
-
- Fares C., Qian J., Davis J.H. Magic angle spinning and static oriented sample NMR studies of the relaxation in the rotating frame of membrane peptides. J. Chem. Phys. 2005;122:194908.
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources

