Electrophysiological characterization of LacY
- PMID: 19383792
- PMCID: PMC2671326
- DOI: 10.1073/pnas.0902471106
Electrophysiological characterization of LacY
Abstract
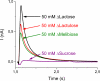
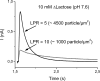
Electrogenic events due to the activity of wild-type lactose permease from Escherichia coli (LacY) were investigated with proteoliposomes containing purified LacY adsorbed on a solid-supported membrane electrode. Downhill sugar/H(+) symport into the proteoliposomes generates transient currents. Studies at different lipid-to-protein ratios and at different pH values, as well as inactivation by N-ethylmaleimide, show that the currents are due specifically to the activity of LacY. From analysis of the currents under different conditions and comparison with biochemical data, it is suggested that the predominant electrogenic event in downhill sugar/H(+) symport is H(+) release. In contrast, LacY mutants Glu-325-->Ala and Cys-154-->Gly, which bind ligand normally, but are severely defective with respect to lactose/H(+) symport, exhibit only a small electrogenic event on addition of LacY-specific substrates, representing 6% of the total charge displacement of the wild-type. This activity is due either to substrate binding per se or to a conformational transition after substrate binding, and is not due to sugar/H(+) symport. We propose that turnover of LacY involves at least 2 electrogenic reactions: (i) a minor electrogenic step that occurs on sugar binding and is due to a conformational transition in LacY; and (ii) a major electrogenic step probably due to cytoplasmic release of H(+) during downhill sugar/H(+) symport, which is the limiting step for this mode of transport.
Conflict of interest statement
The authors declare no conflict of interest.
Figures


Similar articles
-
Delineating electrogenic reactions during lactose/H+ symport.Biochemistry. 2010 Jul 27;49(29):6115-21. doi: 10.1021/bi100492p. Biochemistry. 2010. PMID: 20568736 Free PMC article.
-
Role of protons in sugar binding to LacY.Proc Natl Acad Sci U S A. 2012 Oct 16;109(42):16835-40. doi: 10.1073/pnas.1214890109. Epub 2012 Oct 2. Proc Natl Acad Sci U S A. 2012. PMID: 23033496 Free PMC article.
-
Electrophysiological characterization of uncoupled mutants of LacY.Biochemistry. 2013 Nov 19;52(46):8261-6. doi: 10.1021/bi4013269. Epub 2013 Nov 8. Biochemistry. 2013. PMID: 24152072
-
It takes two to tango: The dance of the permease.J Gen Physiol. 2019 Jul 1;151(7):878-886. doi: 10.1085/jgp.201912377. Epub 2019 May 30. J Gen Physiol. 2019. PMID: 31147449 Free PMC article. Review.
-
The alternating access transport mechanism in LacY.J Membr Biol. 2011 Jan;239(1-2):85-93. doi: 10.1007/s00232-010-9327-5. Epub 2010 Dec 16. J Membr Biol. 2011. PMID: 21161516 Free PMC article. Review.
Cited by
-
Structure of sugar-bound LacY.Proc Natl Acad Sci U S A. 2014 Feb 4;111(5):1784-8. doi: 10.1073/pnas.1324141111. Epub 2014 Jan 22. Proc Natl Acad Sci U S A. 2014. PMID: 24453216 Free PMC article.
-
Evolutionary mix-and-match with MFS transporters II.Proc Natl Acad Sci U S A. 2013 Dec 10;110(50):E4831-8. doi: 10.1073/pnas.1319754110. Epub 2013 Nov 20. Proc Natl Acad Sci U S A. 2013. PMID: 24259711 Free PMC article.
-
Direct observation of electrogenic NH4(+) transport in ammonium transport (Amt) proteins.Proc Natl Acad Sci U S A. 2014 Jul 8;111(27):9995-10000. doi: 10.1073/pnas.1406409111. Epub 2014 Jun 23. Proc Natl Acad Sci U S A. 2014. PMID: 24958855 Free PMC article.
-
Probing of the rates of alternating access in LacY with Trp fluorescence.Proc Natl Acad Sci U S A. 2009 Dec 22;106(51):21561-6. doi: 10.1073/pnas.0911434106. Epub 2009 Dec 3. Proc Natl Acad Sci U S A. 2009. PMID: 19959662 Free PMC article.
-
Delineating electrogenic reactions during lactose/H+ symport.Biochemistry. 2010 Jul 27;49(29):6115-21. doi: 10.1021/bi100492p. Biochemistry. 2010. PMID: 20568736 Free PMC article.
References
-
- Abramson J, et al. Structure and mechanism of the lactose permease of Escherichia coli. Science. 2003;301:610–615. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Molecular Biology Databases

