Ranking differentially expressed genes from Affymetrix gene expression data: methods with reproducibility, sensitivity, and specificity
- PMID: 19386098
- PMCID: PMC2679019
- DOI: 10.1186/1748-7188-4-7
Ranking differentially expressed genes from Affymetrix gene expression data: methods with reproducibility, sensitivity, and specificity
Abstract
Background: To identify differentially expressed genes (DEGs) from microarray data, users of the Affymetrix GeneChip system need to select both a preprocessing algorithm to obtain expression-level measurements and a way of ranking genes to obtain the most plausible candidates. We recently recommended suitable combinations of a preprocessing algorithm and gene ranking method that can be used to identify DEGs with a higher level of sensitivity and specificity. However, in addition to these recommendations, researchers also want to know which combinations enhance reproducibility.
Results: We compared eight conventional methods for ranking genes: weighted average difference (WAD), average difference (AD), fold change (FC), rank products (RP), moderated t statistic (modT), significance analysis of microarrays (samT), shrinkage t statistic (shrinkT), and intensity-based moderated t statistic (ibmT) with six preprocessing algorithms (PLIER, VSN, FARMS, multi-mgMOS (mmgMOS), MBEI, and GCRMA). A total of 36 real experimental datasets was evaluated on the basis of the area under the receiver operating characteristic curve (AUC) as a measure for both sensitivity and specificity. We found that the RP method performed well for VSN-, FARMS-, MBEI-, and GCRMA-preprocessed data, and the WAD method performed well for mmgMOS-preprocessed data. Our analysis of the MicroArray Quality Control (MAQC) project's datasets showed that the FC-based gene ranking methods (WAD, AD, FC, and RP) had a higher level of reproducibility: The percentages of overlapping genes (POGs) across different sites for the FC-based methods were higher overall than those for the t-statistic-based methods (modT, samT, shrinkT, and ibmT). In particular, POG values for WAD were the highest overall among the FC-based methods irrespective of the choice of preprocessing algorithm.
Conclusion: Our results demonstrate that to increase sensitivity, specificity, and reproducibility in microarray analyses, we need to select suitable combinations of preprocessing algorithms and gene ranking methods. We recommend the use of FC-based methods, in particular RP or WAD.
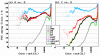
Figures
References
LinkOut - more resources
Full Text Sources
Other Literature Sources

