High-throughput screening of optimal solution conditions for structural biological studies by fluorescence correlation spectroscopy
- PMID: 19388076
- PMCID: PMC2771313
- DOI: 10.1002/pro.92
High-throughput screening of optimal solution conditions for structural biological studies by fluorescence correlation spectroscopy
Abstract
Protein aggregation is an essential molecular event in a wide variety of biological situations, and is a causal factor in several degenerative diseases. The aggregation of proteins also frequently hampers structural biological analyses, such as solution NMR studies. Therefore, precise detection and characterization of protein aggregation are of crucial importance for various research fields. In this study, we demonstrate that fluorescence correlation spectroscopy (FCS) using a single-molecule fluorescence detection system enables the detection of otherwise invisible aggregation of proteins at higher protein concentrations, which are suitable for structural biological experiments, and consumes relatively small amounts of protein over a short measurement time. Furthermore, utilizing FCS, we established a method for high-throughput screening of protein aggregation and optimal solution conditions for structural biological experiments.
Figures

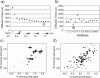
References
-
- Stefani M, Dobson CM. Protein aggregation and aggregate toxicity: new insights into protein folding, misfolding diseases and biological evolution. J Mol Med. 2003;81:678–699. - PubMed
-
- Chayen NE, Shaw Stewart PD, Blow DM. Microbatch crystallization under oil - a new technique allowing many small-volume crystallization trials. J Cryst Growth. 1992;122:176–180.
-
- Bagby S, Tong KI, Liu D, Alattia JR, Ikura M. The button tests: A small scale method using microdialysis cells for assessing protein solubility at concentration suitable fot NMR. J Biomol NMR. 1997;10:279–282. - PubMed
-
- Lepre CA, Moore JM. Microdrop screening: A rapid method to optimize solubent conditions for NMR spectroscopy of proteins. J Biomol NMR. 1998;12:493–499. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources

