Autoprocessing of the Escherichia coli AIDA-I autotransporter: a new mechanism involving acidic residues in the junction region
- PMID: 19398552
- PMCID: PMC2719369
- DOI: 10.1074/jbc.M109.010108
Autoprocessing of the Escherichia coli AIDA-I autotransporter: a new mechanism involving acidic residues in the junction region
Abstract
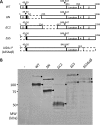
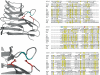
The cleavage of the autotransporter adhesin involved in diffuse adherence (AIDA-I) of Escherichia coli yields a membrane-embedded fragment, AIDAc, and an extracellular fragment, the mature AIDA-I adhesin. The latter remains noncovalently associated with AIDAc but can be released by heat treatment. In this study we determined the mechanism of AIDA-I cleavage. We showed that AIDA-I processing is an autocatalytic event by monitoring the in vitro cleavage of an uncleaved mutant protein isolated from inclusion bodies. Furthermore, by following changes in circular dichroism spectra and protease resistance of the renaturated protein, we showed that the cleavage of the protein is correlated with folding. With site-directed deletions, we showed that the catalytic activity of the protein lies in a region encompassing amino acids between Ala-667 and Thr-953, which includes the conserved junction domain of some autotransporters. With site-directed point mutations, we also found that Asp-878 and Glu-897 are involved in the processing of AIDA-I and that a mutation preserving the acidic side chain of Asp-878 was tolerated, giving evidence that this carboxylic acid group is directly involved in catalysis. Last, we confirmed that cleavage of AIDA-I is intramolecular. Our results unveil a new mechanism of auto-processing in the autotransporter family.
Figures

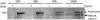





References
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources

