DIRS1-like retrotransposons are widely distributed among Decapoda and are particularly present in hydrothermal vent organisms
- PMID: 19400949
- PMCID: PMC2685390
- DOI: 10.1186/1471-2148-9-86
DIRS1-like retrotransposons are widely distributed among Decapoda and are particularly present in hydrothermal vent organisms
Abstract
Background: Transposable elements are major constituents of eukaryote genomes and have a great impact on genome structure and stability. Considering their mutational abilities, TEs can contribute to the genetic diversity and evolution of organisms. Knowledge of their distribution among several genomes is an essential condition to study their dynamics and to better understand their role in species evolution. DIRS1-like retrotransposons are a particular group of retrotransposons according to their mode of transposition that implies a tyrosine recombinase. To date, they have been described in a restricted number of species in comparison with the LTR retrotransposons. In this paper, we determine the distribution of DIRS1-like elements among 25 decapod species, 10 of them living in hydrothermal vents that correspond to particularly unstable environments.
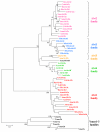
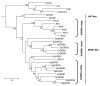
Results: Using PCR approaches, we have identified 15 new DIRS1-like families in 15 diverse decapod species (shrimps, lobsters, crabs and galatheid crabs). Hydrothermal organisms show a particularly great diversity of DIRS1-like elements with 5 families characterized among Alvinocarididae shrimps and 3 in the galatheid crab Munidopsis recta. Phylogenic analyses show that these elements are divergent toward the DIRS1-like families previously described in other crustaceans and arthropods and form a new clade called AlDIRS1. At larger scale, the distribution of DIRS1-like retrotransposons appears more or less patchy depending on the taxa considered. Indeed, a scattered distribution can be observed in the infraorder Brachyura whereas all the species tested in infraorders Caridea and Astacidea harbor some DIRS1-like elements.
Conclusion: Our results lead to nearly double both the number of DIRS1-like elements described to date, and the number of species known to harbor these ones. In this study, we provide the first degenerate primers designed to look specifically for DIRS1-like retrotransposons. They allowed for revealing for the first time a widespread distribution of these elements among a large phylum, here the order Decapoda. They also suggest some peculiar features of these retrotransposons in hydrothermal organisms where a great diversity of elements is already observed. Finally, this paper constitutes the first essential step which allows for considering further studies based on the dynamics of the DIRS1-like retrotransposons among several genomes.
Figures



Similar articles
-
Eukaryote DIRS1-like retrotransposons: an overview.BMC Genomics. 2011 Dec 20;12:621. doi: 10.1186/1471-2164-12-621. BMC Genomics. 2011. PMID: 22185659 Free PMC article. Review.
-
LTR-retrotransposons in R. exoculata and other crustaceans: the outstanding success of GalEa-like copia elements.PLoS One. 2013;8(3):e57675. doi: 10.1371/journal.pone.0057675. Epub 2013 Mar 4. PLoS One. 2013. PMID: 23469217 Free PMC article.
-
GalEa retrotransposons from galatheid squat lobsters (Decapoda, Anomura) define a new clade of Ty1/copia-like elements restricted to aquatic species.Mol Genet Genomics. 2008 Jan;279(1):63-73. doi: 10.1007/s00438-007-0295-0. Epub 2007 Oct 10. Mol Genet Genomics. 2008. PMID: 17929059
-
Divergence history and hydrothermal vent adaptation of decapod crustaceans: A mitogenomic perspective.PLoS One. 2019 Oct 29;14(10):e0224373. doi: 10.1371/journal.pone.0224373. eCollection 2019. PLoS One. 2019. PMID: 31661528 Free PMC article.
-
DIRS-1 and the other tyrosine recombinase retrotransposons.Cytogenet Genome Res. 2005;110(1-4):575-88. doi: 10.1159/000084991. Cytogenet Genome Res. 2005. PMID: 16093711 Review.
Cited by
-
Eukaryote DIRS1-like retrotransposons: an overview.BMC Genomics. 2011 Dec 20;12:621. doi: 10.1186/1471-2164-12-621. BMC Genomics. 2011. PMID: 22185659 Free PMC article. Review.
-
Filamentous ascomycete genomes provide insights into Copia retrotransposon diversity in fungi.BMC Genomics. 2017 May 25;18(1):410. doi: 10.1186/s12864-017-3795-2. BMC Genomics. 2017. PMID: 28545447 Free PMC article.
-
Mobilization of retrotransposons as a cause of chromosomal diversification and rapid speciation: the case for the Antarctic teleost genus Trematomus.BMC Genomics. 2018 May 9;19(1):339. doi: 10.1186/s12864-018-4714-x. BMC Genomics. 2018. PMID: 29739320 Free PMC article.
-
LTR-retrotransposons in R. exoculata and other crustaceans: the outstanding success of GalEa-like copia elements.PLoS One. 2013;8(3):e57675. doi: 10.1371/journal.pone.0057675. Epub 2013 Mar 4. PLoS One. 2013. PMID: 23469217 Free PMC article.
-
The evolution of tyrosine-recombinase elements in Nematoda.PLoS One. 2014 Sep 8;9(9):e106630. doi: 10.1371/journal.pone.0106630. eCollection 2014. PLoS One. 2014. PMID: 25197791 Free PMC article.
References
-
- Nyholm SV, Robidart J, Girguis PR. Coupling metabolite flux to transcriptomics: insights into the molecular mechanisms underlying primary productivity by the hydrothermal vent tubeworm Ridgeia piscesae. Biol Bull. 2008;214:255–265. - PubMed
-
- Sarradin P, Caprais J, Riso R, Kerouel R, Aminot A. Chemical environment of the hydrothermal mussel communities in the Lucky Strike and Menez Gwen vent fields, Mid Atlantic Ridge. Cahiers de biologie marine. 1999;40:93–104.
-
- Shank TM, Fornari DJ, Von Damm KL, Lilley MD, Haymon RM, Lutz RA. Temporal and spatial patterns of biological community development at nascent deep-sea hydrothermal vents (9°50'N, East Pacific Rise) Deep Sea Research Part II: Topical Studies in Oceanography. 1998;45:465–515. doi: 10.1016/S0967-0645(97)00089-1. - DOI
MeSH terms
Substances
LinkOut - more resources
Full Text Sources

