Gene regulatory network inference using out of equilibrium statistical mechanics
- PMID: 19404429
- PMCID: PMC2639940
- DOI: 10.2976/1.2957743
Gene regulatory network inference using out of equilibrium statistical mechanics
Abstract
Spatiotemporal control of gene expression is fundamental to multicellular life. Despite prodigious efforts, the encoding of gene expression regulation in eukaryotes is not understood. Gene expression analyses nourish the hope to reverse engineer effector-target gene networks using inference techniques. Inference from noisy and circumstantial data relies on using robust models with few parameters for the underlying mechanisms. However, a systematic path to gene regulatory network reverse engineering from functional genomics data is still impeded by fundamental problems. Recently, Johannes Berg from the Theoretical Physics Institute of Cologne University has made two remarkable contributions that significantly advance the gene regulatory network inference problem. Berg, who uses gene expression data from yeast, has demonstrated a nonequilibrium regime for mRNA concentration dynamics and was able to map the gene regulatory process upon simple stochastic systems driven out of equilibrium. The impact of his demonstration is twofold, affecting both the understanding of the operational constraints under which transcription occurs and the capacity to extract relevant information from highly time-resolved expression data. Berg has used his observation to predict target genes of selected transcription factors, and thereby, in principle, demonstrated applicability of his out of equilibrium statistical mechanics approach to the gene network inference problem.
Figures
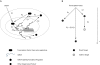

References
-
- Adams, D (1979). The Hitchhiker’s guide to the galaxy, Pan Books, London.
-
- Barrera, L O, and Ren, B, (2006). “The transcriptional regulatory code of eukaryotic cells—insights from genome-wide analysis of chromatin organization and transcription factor binding.” Curr. Opin. Cell Biol. 18, 291–2980. - PubMed
-
- Benecke, A (2003). “Genomic plasticity and information processing by transcription coregulators.” ComPlexUs 1, 65–76.
-
- Benecke, A (2006). “Chromatin code, local non-equilibrium dynamics, and the emergence of transcription regulatory programs.” Eur. Phys. J. E 19, 379–384. - PubMed
LinkOut - more resources
Full Text Sources
Molecular Biology Databases
