Strategies and issues in the detection of pathway enrichment in genome-wide association studies
- PMID: 19408013
- PMCID: PMC2865249
- DOI: 10.1007/s00439-009-0676-z
Strategies and issues in the detection of pathway enrichment in genome-wide association studies
Abstract
A fundamental question in human genetics is the degree to which the polygenic character of complex traits derives from polymorphism in genes with similar or with dissimilar functions. The many genome-wide association studies now being performed offer an opportunity to investigate this, and although early attempts are emerging, new tools and modeling strategies still need to be developed and deployed. Towards this goal, we implemented a new algorithm to facilitate the transition from genetic marker lists (principally those generated by PLINK) to pathway analyses of representational gene sets in either threshold or threshold-free downstream applications (e.g. DAVID, GSEA-P, and Ingenuity Pathway Analysis). This was applied to several large genome-wide association studies covering diverse human traits that included type 2 diabetes, Crohn's disease, and plasma lipid levels. Validation of this approach was obtained for plasma HDL levels, where functional categories related to lipid metabolism emerged as the most significant in two independent studies. From analyses of these samples, we highlight and address numerous issues related to this strategy, including appropriate gene based correction statistics, the utility of imputed versus non-imputed marker sets, and the apparent enrichment of pathways due solely to the positional clustering of functionally related genes. The latter in particular emphasizes the importance of studies that directly tie genetic variation to functional characteristics of specific genes. The software freely provided that we have called ProxyGeneLD may resolve an important bottleneck in pathway-based analyses of genome-wide association data. This has allowed us to identify at least one replicable case of pathway enrichment but also to highlight functional gene clustering as a potentially serious problem that may lead to spurious pathway findings if not corrected.
Conflict of interest statement
The authors declare no conflict of interest.
Figures
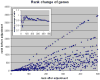
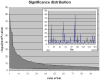
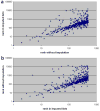
References
-
- Ashburner M, Ball CA, Blake JA, Botstein D, Butler H, Cherry JM, Davis AP, Dolinski K, Dwight SS, Eppig JT, Harris MA, Hill DP, Issel-Tarver L, Kasarskis A, Lewis S, Matese JC, Richardson JE, Ringwald M, Rubin GM, Sherlock G. Gene ontology: tool for the unification of biology. The Gene Ontology Consortium. Nat Genet. 2000;25:25–9. - PMC - PubMed
-
- Askland K, Read C, Moore J. Pathways-based analyses of whole-genome association study data in bipolar disorder reveal genes mediating ion channel activity and synaptic neurotransmission. Hum Genet. 2009;125:63–79. - PubMed
-
- Aulchenko YS, Ripatti S, Lindqvist I, Boomsma D, Heid IM, Pramstaller PP, Penninx BW, Janssens AC, Wilson JF, Spector T, Martin NG, Pedersen NL, Kyvik KO, Kaprio J, Hofman A, Freimer NB, Jarvelin MR, Gyllensten U, Campbell H, Rudan I, Johansson A, Marroni F, Hayward C, Vitart V, Jonasson I, Pattaro C, Wright A, Hastie N, Pichler I, Hicks AA, Falchi M, Willemsen G, Hottenga JJ, de Geus EJ, Montgomery GW, Whitfield J, Magnusson P, Saharinen J, Perola M, Silander K, Isaacs A, Sijbrands EJ, Uitterlinden AG, Witteman JC, Oostra BA, Elliott P, Ruokonen A, Sabatti C, Gieger C, Meitinger T, Kronenberg F, Doring A, Wichmann HE, Smit JH, McCarthy MI, van Duijn CM, Peltonen L. Loci influencing lipid levels and coronary heart disease risk in 16 European population cohorts. Nat Genet. 2009;41:47–55. - PMC - PubMed
-
- Baranzini SE, Wang J, Gibson RA, Galwey N, Naegelin Y, Barkhof F, Radue EW, Lindberg RL, Uitdehaag B, Johnson MR, Angelakopoulou A, Hall L, Richardson JC, Prinjha RK, Gass A, Geurts JJ, Kragt J, Sombekke M, Vrenken H, Qualley P, Lincoln RR, Gomez R, Caillier SJ, George MF, Mousavi H, Guerrero R, Okuda DT, Cree BA, Green A, Waubant E, Goodin DS, Pelletier D, Matthews PM, Hauser SL, Kappos L, Polman CH, Oksenberg JR. Genome-wide association analysis of susceptibility and clinical phenotype in multiple sclerosis. Hum Mol Genet. 2009;18:767–78. - PMC - PubMed
-
- Barrett JC, Hansoul S, Nicolae DL, Cho JH, Duerr RH, Rioux JD, Brant SR, Silverberg MS, Taylor KD, Barmada MM, Bitton A, Dassopoulos T, Datta LW, Green T, Griffiths AM, Kistner EO, Murtha MT, Regueiro MD, Rotter JI, Schumm LP, Steinhart AH, Targan SR, Xavier RJ, Libioulle C, Sandor C, Lathrop M, Belaiche J, Dewit O, Gut I, Heath S, Laukens D, Mni M, Rutgeerts P, Van Gossum A, Zelenika D, Franchimont D, Hugot JP, de Vos M, Vermeire S, Louis E, Cardon LR, Anderson CA, Drummond H, Nimmo E, Ahmad T, Prescott NJ, Onnie CM, Fisher SA, Marchini J, Ghori J, Bumpstead S, Gwilliam R, Tremelling M, Deloukas P, Mansfield J, Jewell D, Satsangi J, Mathew CG, Parkes M, Georges M, Daly MJ. Genome-wide association defines more than 30 distinct susceptibility loci for Crohn’s disease. Nat Genet. 2008;40:955–62. - PMC - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Miscellaneous

