NKAP is a transcriptional repressor of notch signaling and is required for T cell development
- PMID: 19409814
- PMCID: PMC2777751
- DOI: 10.1016/j.immuni.2009.02.011
NKAP is a transcriptional repressor of notch signaling and is required for T cell development
Abstract
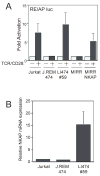
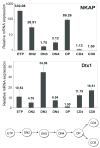
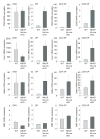
T cell development depends on the coordinated interplay between receptor signaling and transcriptional regulation. Through a genetic complementation screen a transcriptional repressor, NKAP, was identified. NKAP associated with the histone deacetylase HDAC3 and was shown to be part of a DNA-binding complex, as demonstrated by chromatin immunoprecipitation. NKAP also associated with the Notch corepressor complex. The expression of NKAP during T cell development inversely correlated with the expression of Notch target genes, implying that NKAP may modulate Notch-mediated transcription. To examine the function of NKAP in T cell development, we ablated NKAP by Lck(cre). Loss of NKAP blocked development of alphabeta but not gammadelta T cells, and Nkap(fl/o)Lck(cre) DP T cells expressed 8- to 20-fold higher amounts of Hes1, Deltex1, and CD25 mRNA. Thus, NKAP functions as a transcriptional repressor, acting on Notch target genes, and is required for alphabeta T cell development.
Conflict of interest statement
The authors state that they have no financial conflict of interest.
Figures




References
-
- Abraham RT, Weiss A. Jurkat T cells and development of the T-cell receptor signalling paradigm. Nat Rev Immunol. 2004;4:301–308. - PubMed
-
- Chen D, Li Z, Yang Q, Zhang J, Zhai Z, Shu HB. Identification of a nuclear protein that promotes NF-κB activation. Biochem Biophys Res Commun. 2003;310:720–724. - PubMed
-
- Dohda T, Maljukova A, Liu L, Heyman M, Grander D, Brodin D, Sangfelt O, Lendahl U. Notch signaling induces Skp2 expression and promotes reduction of p27Kip1 in T-cell acute lymphoblastic leukemia cell lines. Exp Cell Res. 2007;313:3141–3142. - PubMed
-
- Ellisen LW, Bird J, West DC, Soreng AL, Reynolds TC, Smith SD, Sklar J. TAN-1, the human homolog of the Drosophila notch gene, is broken by chromosomal translocations in T lymphoblastic neoplasms. Cell. 1991;66:649–661. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Molecular Biology Databases
Miscellaneous

