Analysis of post-transcriptional regulations by a functional, integrated, and quantitative method
- PMID: 19411282
- PMCID: PMC2722765
- DOI: 10.1074/mcp.M800503-MCP200
Analysis of post-transcriptional regulations by a functional, integrated, and quantitative method
Abstract
In the past 10 years, transcriptome and proteome analyses have provided valuable data on global gene expression and cell functional networks. However, when integrated,these analyses revealed partial correlations between mRNA expression levels and protein abundance thus suggesting that post-transcriptional regulations may be in part responsible for this discrepancy. In the present work, we report the development of a functional, integrated, and quantitative method to measure post-transcriptional regulations that we named FunREG. This method enables (i) quantitative measure of post-transcriptional regulations mediated by selected 3-untranslated regions and exogenous small interfering-RNA or micro-RNAs and (ii) comparison of these regulatory processes in physiologically relevant systems (e.g. cancer versus primary untransformed cells). We applied FunREG to the study of liver cancer, and we demonstrate for the first time the differential regulatory mechanisms controlling gene expression at a post-transcriptional level in normal and tumoral hepatic cells. As an example, translation efficiency mediated by heparin-binding epidermal growth factor 3-untranslated region was increased 3-fold in liver cancer cells compared with normal hepatocytes, whereas stability of an mRNA containing a portion of Cyclin D1 3-untranslated region was increased more than 2-fold in HepG2 cells compared with normal hepatocytes. Consequently we believe that the method presented herein may become an important tool in fundamental and medical research. This approach is convenient and easy to perform, accessible to any investigator, and should be adaptable to a large number of cell type, functional and chemical screens, as well as genome scale analyses. Finally FunREG may represent a helpful tool to reconcile transcriptome and proteome data.
Figures




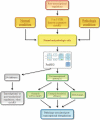

References
-
- Gallardo K., Firnhaber C., Zuber H., Héricher D., Belghazi M., Henry C., Küster H., Thompson R. ( 2007) A combined proteome and transcriptome analysis of developing Medicago truncatula seeds: evidence for metabolic specialization of maternal and filial tissues. Mol. Cell. Proteomics 6, 2165– 2179 - PubMed
-
- Williamson A. J., Smith D. L., Blinco D., Unwin R. D., Pearson S., Wilson C., Miller C., Lancashire L., Lacaud G., Kouskoff V., Whetton A. D. ( 2008) Quantitative proteomics analysis demonstrates post-transcriptional regulation of embryonic stem cell differentiation to hematopoiesis. Mol. Cell. Proteomics 7, 459– 472 - PubMed
-
- Beyer A., Hollunder J., Nasheuer H. P., Wilhelm T. ( 2004) Post-transcriptional expression regulation in the yeast Saccharomyces cerevisiae on a genomic scale. Mol. Cell. Proteomics 3, 1083– 1092 - PubMed
-
- Minagawa H., Honda M., Miyazaki K., Tabuse Y., Teramoto R., Yamashita T., Nishino R., Takatori H., Ueda T., Kamijo K., Kaneko S. ( 2008) Comparative proteomic and transcriptomic profiling of the human hepatocellular carcinoma. Biochem. Biophys. Res. Commun. 366, 186– 192 - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Research Materials
Miscellaneous

