Novel micropatterned cardiac cell cultures with realistic ventricular microstructure
- PMID: 19413993
- PMCID: PMC3297758
- DOI: 10.1016/j.bpj.2009.02.019
Novel micropatterned cardiac cell cultures with realistic ventricular microstructure
Abstract
Systematic studies of cardiac structure-function relationships to date have been hindered by the intrinsic complexity and variability of in vivo and ex vivo model systems. Thus, we set out to develop a reproducible cell culture system that can accurately replicate the realistic microstructure of native cardiac tissues. Using cell micropatterning techniques, we aligned cultured cardiomyocytes at micro- and macroscopic spatial scales to follow local directions of cardiac fibers in murine ventricular cross sections, as measured by high-resolution diffusion tensor magnetic resonance imaging. To elucidate the roles of ventricular tissue microstructure in macroscopic impulse conduction, we optically mapped membrane potentials in micropatterned cardiac cultures with realistic tissue boundaries and natural cell orientation, cardiac cultures with realistic tissue boundaries but random cell orientation, and standard isotropic monolayers. At 2 Hz pacing, both microscopic changes in cell orientation and ventricular tissue boundaries independently and synergistically increased the spatial dispersion of conduction velocity, but not the action potential duration. The realistic variations in intramural microstructure created unique spatial signatures in micro- and macroscopic impulse propagation within ventricular cross-section cultures. This novel in vitro model system is expected to help bridge the existing gap between experimental structure-function studies in standard cardiac monolayers and intact heart tissues.
Figures





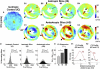


References
-
- de Bakker J.M., van Capelle F.J., Janse M.J., Tasseron S., Vermeulen J.T. Slow conduction in the infarcted human heart. ‘Zigzag’ course of activation. Circulation. 1993;88:915–926. - PubMed
-
- Valderrabano M., Lee M.H., Ohara T., Lai A.C., Fishbein M.C. Dynamics of intramural and transmural reentry during ventricular fibrillation in isolated swine ventricles. Circ. Res. 2001;88:839–848. - PubMed
-
- Muthappan P., Calkins H. Arrhythmogenic right ventricular dysplasia. Prog. Cardiovasc. Dis. 2008;51:31–43. - PubMed
-
- Schalij M.J., Boersma L., Huijberts M., Allessie M.A. Anisotropic reentry in a perfused 2-dimensional layer of rabbit ventricular myocardium. Circulation. 2000;102:2650–2658. - PubMed
Publication types
MeSH terms
LinkOut - more resources
Full Text Sources
Other Literature Sources
Miscellaneous

