Development of a reverse transcription multiplex real-time PCR for the detection and genotyping of classical swine fever virus
- PMID: 19414034
- PMCID: PMC7112934
- DOI: 10.1016/j.jviromet.2009.04.029
Development of a reverse transcription multiplex real-time PCR for the detection and genotyping of classical swine fever virus
Abstract
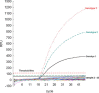
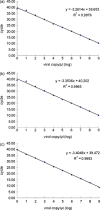
A reverse transcription multiplex real-time PCR (RT-MRT-PCR) was developed for rapid detection and genotyping of classical swine fever virus (CSFV). The universal primers and specific TaqMan probes for each of the three genotypes, genotypes 1, 2, and 3, were designed within the 3'-UTR of the CSFV. Non-CSFV swine virus and clinical samples from specific pathogen-free (SPF) pigs were both demonstrated to be CSFV-negative by RT-MRT-PCR. The diagnostic sensitivity of RT-MRT-PCR was determined to be 1 viral copy/microl for each genotype of standard plasmid. For the analytical sensitivity experiment, 100 samples of 14 CSFV genotype 1 strains and 86 samples from CSFV outbreak farms were all detected as CSFV-positive by RT-MRT-PCR, and the genotype results were consistent with the results of sequencing from a previous study. The intra-assay and inter-assay variations of RT-MRT-PCR were below 3% in all experiments. The sensitivity of RT-MRT-PCR was the same as the reverse transcription nested PCR (RT-nPCR) and higher than reverse transcription PCR (RT-PCR) and viral isolation from clinical samples. This assay was used further to evaluate the duration of viremia of wild-type CSFV in vaccinated exposed pigs. The results indicated that pigs vaccinated with the E2 subunit vaccine had longer viremia than pigs given the C-strain vaccine, which is compatible with the findings of previous studies. Thus, the new RT-MRT-PCR is a rapid, reproducible, sensitive, and specific genotyping tool for CSFV detection.
Figures
References
-
- Baxi M., McRae D., Boxi S., Greiser-Wike I., Vilcek S., Amoako K., Deregt D. A one-step multiplex real-time RT-PCR for detection and typing of bovine viral diarrhea viruses. Vet. Microbiol. 2006;116:37–44. - PubMed
-
- Biagetti M., Greiser-Wilke I., Rutili D. Molecular epidemiology of classical swine fever in Italy. Vet. Microbiol. 2001;83:205–215. - PubMed
-
- Blacksell S.D., Khounsy S., Boyle D.B., Gleeson L.J., Westbury H.A., Mackenzie J.S. Genetic typing of classical swine fever viruses from Lao PDR by analysis of the 5′ non-coding region. Virus Genes. 2005;31:349–355. - PubMed
-
- Depner K., Hoffmann B., Beer M. Evaluation of real-time RT-PCR assay for the routine intra vitam diagnosis of classical swine fever. Vet. Microbiol. 2007;121:338–343. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources

