Phylogenetically distinct Staphylococcus aureus lineage prevalent among indigenous communities in northern Australia
- PMID: 19420161
- PMCID: PMC2708510
- DOI: 10.1128/JCM.00122-09
Phylogenetically distinct Staphylococcus aureus lineage prevalent among indigenous communities in northern Australia
Abstract
The aim was to determine the evolutionary position of the Staphylococcus aureus clonal complex 75 (CC75) that is prevalent in tropical northern Australia. Sequencing of gap, rpoB, sodA, tuf, and hsp60 and the multilocus sequence typing loci revealed a clear separation between conventional S. aureus and CC75 and significant diversity within CC75.
Figures
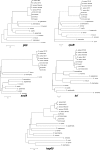
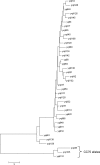
References
-
- Euzéby, J. P. 1997. List of bacterial names with standing in nomenclature: a folder available on the Internet. Int. J. Syst. Bacteriol. 47590-592. - PubMed
Publication types
MeSH terms
Substances
Associated data
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
LinkOut - more resources
Full Text Sources
Medical
Research Materials
Miscellaneous

