The functional effect of pathogenic mutations in Rab escort protein 1
- PMID: 19427510
- PMCID: PMC2680797
- DOI: 10.1016/j.mrfmmm.2009.02.015
The functional effect of pathogenic mutations in Rab escort protein 1
Abstract
Choroideremia (CHM) is a chorioretinal degeneration with an X-linked pattern of inheritance. Affected males experience progressive atrophy of the choroid, retinal pigment epithelium and retina leading to eventual blindness. The CHM gene encodes Rab escort protein 1 (REP-1). REP-1 is involved in trafficking of Rab proteins in the cell. To date, the majority of reported mutations in the CHM gene cause a complete loss of REP-1 function. Here we report pathogenic mutations: a novel missense mutation, L550P; a truncation c.1542T>A, STOP; and two deletions (c.525_526delAG and c.1646delC) in the CHM gene and their phenotypic effect. To analyze the effect of mutations, the 3D structure of human REP-1 and the proteins associated with REP-1 function were modeled using sequence homology with rat proteins. In silico analysis of the missense mutation L550P suggests that the proline residue at position 550 destabilizes the beta-structural elements, and the REP-1 tertiary structure. Truncation and deletion mutants are associated with a partial or total loss of the REP-1 essential activity and protein-protein interactions as predicted by the analysis of the structure and stability of these protein products. The presumptive loss of protein was confirmed by Western Blot analysis of protein from mononuclear cells and fibroblasts (FB) from CHM patients.
Figures
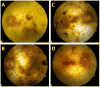



References
-
- Seabra MC, Brown MS, Slaughter CA, Sudhof TC, Goldstein JL. Purification of component A of Rab geranylgeranyl transferase: possible identity with the choroideremia gene product. Cell. 1992;70:1049–1057. - PubMed
-
- Seabra MC, Brown MS, Goldstein JL. Retinal degeneration in choroideremia: deficiency of rab geranylgeranyl transferase. Science. 1993;259:377–381. - PubMed
-
- Andres DA, Seabra MC, Brown MS, Armstrong SA, Smeland TE, Cremers FP, Goldstein JL. cDNA cloning of component A of Rab geranylgeranyl transferase and demonstration of its role as a Rab escort protein. Cell. 1993;73:1091–1099. - PubMed
-
- MacDonald IM, Mah DY, Ho YK, Lewis RA, Seabra MC. A practical diagnostic test for choroideremia. Ophthalmology. 1998;105:1637–1640. - PubMed
-
- MacDonald IM, Sereda C, McTaggart K, Mah D. Choroideremia gene testing. Expert Rev Mol Diagn. 2004;4:478–484. - PubMed
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Molecular Biology Databases
Research Materials

