IRF-1 and p65 mediate upregulation of constitutive HLA-A antigen expression by hepatocellular carcinoma cells
- PMID: 19428110
- PMCID: PMC3426235
- DOI: 10.1016/j.molimm.2009.03.001
IRF-1 and p65 mediate upregulation of constitutive HLA-A antigen expression by hepatocellular carcinoma cells
Abstract
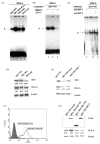
Malignant transformation of hepatocytes is frequently associated with upregulation of HLA-A expression. Currently there is no information available regarding the mechanisms underlying this phenotypic change. We investigated HLA-A expression in 165 paraffin embedded tissues and 21 fresh tissues from liver cancer patients. Utilizing truncated HLA-A promoter-reporter constructs and gel-shift assay we had identified the regulatory elements and transcription factors required for HLA-A upregulation. 54% of the paraffin embedded tissues showed increased HLA-A expression in their cancerous part. 43% of the fresh liver cancer tissues had increased HLA-A complex expression with the HLA-A heavy chain gene demonstrating the highest level of upregulation (62%). Enhanced HLA-A expression in the liver cell lines QGY7701 and BEL7402 was found to be mediated by binding of interferon regulatory factor 1 (IRF-1) to interferon stimulated response element, and of nuclear transcription factor p65 binding to enhancer A element in the HLA-A promoter of these cell lines. The in vivo relevance of these findings was indicated by the association of the enhanced expression of IRF-1 and accumulation of nuclear p65 with HLA-A upregulation in 8 of the 21 liver cancer lesions investigated. Our results indicated that HLA-A upregulation in liver cancer was mediated by both increased nuclear aggregation of transcription factor p65 and upregulation of transcription factor IRF-1.
Figures



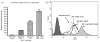

Similar articles
-
Histone deacetylase-1 is enriched at the platelet-derived growth factor-D promoter in response to interleukin-1beta and forms a cytokine-inducible gene-silencing complex with NF-kappab p65 and interferon regulatory factor-1.J Biol Chem. 2009 Dec 11;284(50):35101-12. doi: 10.1074/jbc.M109.061903. Epub 2009 Oct 20. J Biol Chem. 2009. Retraction in: J Biol Chem. 2010 Jul 9;285(28):21902. doi: 10.1074/jbc.a209.061903. PMID: 19843519 Free PMC article. Retracted.
-
CD40 induces antigen transporter and immunoproteasome gene expression in carcinomas via the coordinated action of NF-kappaB and of NF-kappaB-mediated de novo synthesis of IRF-1.Mol Cell Biol. 2008 Oct;28(20):6208-22. doi: 10.1128/MCB.00611-08. Epub 2008 Aug 11. Mol Cell Biol. 2008. PMID: 18694960 Free PMC article.
-
Leptin augments recruitment of IRF-1 and CREB to thrombospondin-1 gene promoter in vascular smooth muscle cells in vitro.Am J Physiol Cell Physiol. 2016 Aug 1;311(2):C212-24. doi: 10.1152/ajpcell.00068.2016. Epub 2016 Jun 8. Am J Physiol Cell Physiol. 2016. PMID: 27281481
-
MicroRNA-301a (miR-301a) is induced in hepatocellular carcinoma (HCC) and down- regulates the expression of interferon regulatory factor-1.Biochem Biophys Res Commun. 2020 Apr 2;524(2):273-279. doi: 10.1016/j.bbrc.2020.01.034. Epub 2020 Jan 25. Biochem Biophys Res Commun. 2020. PMID: 31987500 Free PMC article.
-
An interferon-sensitive response element is involved in constitutive caspase-8 gene expression in neuroblastoma cells.Int J Cancer. 2007 Jan 1;120(1):39-47. doi: 10.1002/ijc.22173. Int J Cancer. 2007. PMID: 17036321
Cited by
-
Maintenance of Hypoimmunogenic Features via Regulation of Endogenous Antigen Processing and Presentation Machinery.Front Bioeng Biotechnol. 2022 Jul 22;10:936584. doi: 10.3389/fbioe.2022.936584. eCollection 2022. Front Bioeng Biotechnol. 2022. PMID: 36032723 Free PMC article.
-
Frequent HLA class I alterations in human prostate cancer: molecular mechanisms and clinical relevance.Cancer Immunol Immunother. 2016 Jan;65(1):47-59. doi: 10.1007/s00262-015-1774-5. Epub 2015 Nov 26. Cancer Immunol Immunother. 2016. PMID: 26611618 Free PMC article.
-
Identification of an HLA-A*24:02-restricted α-fetoprotein signal peptide-derived antigen and its specific T-cell receptor for T-cell immunotherapy.Immunology. 2020 Apr;159(4):384-392. doi: 10.1111/imm.13168. Epub 2020 Jan 10. Immunology. 2020. PMID: 31849039 Free PMC article.
-
Selection of a Clinical Lead TCR Targeting Alpha-Fetoprotein-Positive Liver Cancer Based on a Balance of Risk and Benefit.Front Immunol. 2020 Apr 27;11:623. doi: 10.3389/fimmu.2020.00623. eCollection 2020. Front Immunol. 2020. PMID: 32425926 Free PMC article. Clinical Trial.
-
Identification of IRF1 as critical dual regulator of Smac mimetic-induced apoptosis and inflammatory cytokine response.Cell Death Dis. 2014 Dec 11;5(12):e1562. doi: 10.1038/cddis.2014.498. Cell Death Dis. 2014. PMID: 25501823 Free PMC article.
References
-
- Bi YC, Zhang JQ, Hong ZH, Huang J, Chen JG, Wang XH, Ferrone S, Xie W. Association of HLA class I antigen expression in human carcinoma with improved prognosis. Tissue Antigens. 2002;59s2:105–106.
-
- Blanchet O, Gazin C, L’Haridon M, Tatari Z, Degos L, Sigaux F, Paul P. Multiple nuclear factors bind to novel positive and negative regulatory elements upstream of the human MHC class I gene HLA-A11. Int Immunol. 1994;6:1485–1496. - PubMed
-
- Bodmer WF, Browning MJ, Krausa P, Rowan A, Bicknell DC, Bodmer JG. Tumor escape from immune response by variation in HLA expression and other mechanisms. Ann N Y Acad Sci. 1993;690:42–49. - PubMed
-
- Bouker KB, Skaar TC, Riggins RB, Harburger DS, Fernandez DR, Zwart A, Wang A, Clarke R. Interferon regulatory factor-1 (IRF-1) exhibits tumor suppressor activities in breast cancer associated with caspase activation and induction of apoptosis. Carcinogenesis. 2005;26:1527–1535. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Medical
Molecular Biology Databases
Research Materials

