Community genomic and proteomic analyses of chemoautotrophic iron-oxidizing "Leptospirillum rubarum" (Group II) and "Leptospirillum ferrodiazotrophum" (Group III) bacteria in acid mine drainage biofilms
- PMID: 19429552
- PMCID: PMC2704813
- DOI: 10.1128/AEM.02943-08
Community genomic and proteomic analyses of chemoautotrophic iron-oxidizing "Leptospirillum rubarum" (Group II) and "Leptospirillum ferrodiazotrophum" (Group III) bacteria in acid mine drainage biofilms
Abstract
We analyzed near-complete population (composite) genomic sequences for coexisting acidophilic iron-oxidizing Leptospirillum group II and III bacteria (phylum Nitrospirae) and an extrachromosomal plasmid from a Richmond Mine, Iron Mountain, CA, acid mine drainage biofilm. Community proteomic analysis of the genomically characterized sample and two other biofilms identified 64.6% and 44.9% of the predicted proteins of Leptospirillum groups II and III, respectively, and 20% of the predicted plasmid proteins. The bacteria share 92% 16S rRNA gene sequence identity and >60% of their genes, including integrated plasmid-like regions. The extrachromosomal plasmid carries conjugation genes with detectable sequence similarity to genes in the integrated conjugative plasmid, but only those on the extrachromosomal element were identified by proteomics. Both bacterial groups have genes for community-essential functions, including carbon fixation and biosynthesis of vitamins, fatty acids, and biopolymers (including cellulose); proteomic analyses reveal these activities. Both Leptospirillum types have multiple pathways for osmotic protection. Although both are motile, signal transduction and methyl-accepting chemotaxis proteins are more abundant in Leptospirillum group III, consistent with its distribution in gradients within biofilms. Interestingly, Leptospirillum group II uses a methyl-dependent and Leptospirillum group III a methyl-independent response pathway. Although only Leptospirillum group III can fix nitrogen, these proteins were not identified by proteomics. The abundances of core proteins are similar in all communities, but the abundance levels of unique and shared proteins of unknown function vary. Some proteins unique to one organism were highly expressed and may be key to the functional and ecological differentiation of Leptospirillum groups II and III.
Figures

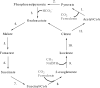



References
-
- Aguilar, P. S., and D. de Mendoza. 2006. Control of fatty acid desaturation: a mechanism conserved from bacteria to humans. Mol. Microbiol. 62:1507-1514. - PubMed
-
- Altschul, S. F., W. Gish, W. Miller, E. W. Myers, and D. J. Lipman. 1990. Basic local alignment search tool. J. Mol. Biol. 215:403-410. - PubMed
-
- Andersson, A. F., and J. F. Banfield. 2008. Virus population dynamics and acquired virus resistance in natural microbial communities. Science 320:1047-1050. - PubMed
-
- Aoshima, M., and Y. Igarashi. 2006. A novel oxalosuccinate-forming enzyme involved in the reductive carboxylation of 2-oxoglutarate in Hydrogenobacter thermophilus TK-6. Mol. Microbiol. 62:748-759. - PubMed
Publication types
MeSH terms
Substances
Associated data
- Actions
- Actions
LinkOut - more resources
Full Text Sources
Molecular Biology Databases

