Two-step assembly dynamics of the Bacillus subtilis divisome
- PMID: 19429628
- PMCID: PMC2698510
- DOI: 10.1128/JB.01758-08
Two-step assembly dynamics of the Bacillus subtilis divisome
Abstract
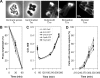
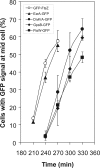
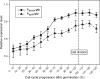
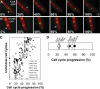
Cell division in bacteria is carried out by about a dozen proteins which assemble at midcell and form a complex known as the divisome. To study the dynamics and temporal hierarchy of divisome assembly in Bacillus subtilis, we have examined the in vivo localization pattern of a set of division proteins fused to green fluorescent protein in germinating spores and vegetative cells. Using time series and time-lapse microscopy, we show that the FtsZ ring assembles early and concomitantly with FtsA, ZapA, and EzrA. After a time delay of at least 20% of the cell cycle, a second set of division proteins, including GpsB, FtsL, DivIB, FtsW, Pbp2B, and DivIVA, are recruited to midcell. Together, our data provide in vivo evidence for two-step assembly of the divisome. Interestingly, overproduction of FtsZ advances the temporal assembly of EzrA but not of DivIVA, suggesting that a signal different from that of FtsZ polymerization drives the assembly of late divisome proteins. Microarray analysis shows that FtsZ depletion or overexpression does not significantly alter the transcription of division genes, supporting the hypothesis that cell division in B. subtilis is mainly regulated at the posttranscriptional level.
Figures


References
-
- Aarsman, M. E., A. Piette, C. Fraipont, T. M. Vinkenvleugel, M. Nguyen-Disteche, and T. den Blaauwen. 2005. Maturation of the Escherichia coli divisome occurs in two steps. Mol. Microbiol. 551631-1645. - PubMed
-
- Barák, I., and A. J. Wilkinson. 2007. Division site recognition in Escherichia coli and Bacillus subtilis. FEMS Microbiol. Rev. 31311-326. - PubMed
-
- Bi, E. F., and J. Lutkenhaus. 1991. FtsZ ring structure associated with division in Escherichia coli. Nature 354161-164. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Molecular Biology Databases

