Development-dependent changes in the tight DNA-protein complexes of barley on chromosome and gene level
- PMID: 19435519
- PMCID: PMC2694405
- DOI: 10.1186/1471-2229-9-56
Development-dependent changes in the tight DNA-protein complexes of barley on chromosome and gene level
Abstract
Background: The tightly bound to DNA proteins (TBPs) is a protein group that remains attached to DNA with covalent or non-covalent bonds after its deproteinisation. The functional role of this group is as yet not completely understood. The main goal of this study was to evaluate tissue specific changes in the TBP distribution in barley genes and chromosomes in different phases of shoot and seed development. We have: 1. investigated the TBP distribution along Amy32b and Bmy1 genes encoding low pI alpha-amylase A and endosperm specific beta-amylase correspondingly using oligonucleotide DNA arrays; 2. characterized the polypeptide spectrum of TBP and proteins with affinity to TBP-associated DNA; 3. localized the distribution of DNA complexes with TBP (TBP-DNA) on barley 1H and 7H chromosomes using mapped markers; 4. compared the chromosomal distribution of TBP-DNA complexes to the distribution of the nuclear matrix attachment sites.
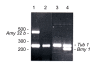
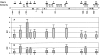
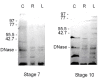
Results: In the Amy32b gene transition from watery ripe to the milky ripeness stage of seed development was followed by the decrease of TBP binding along the whole gene, especially in the promoter region and intron II. Expression of the Bmy1 gene coupled to ripening was followed by release of the exon III and intron III sequences from complexes with TBPs. Marker analysis revealed changes in the association of chromosome 1H and 7H sites with TBPs between first leaf and coleoptile and at Zadoks 07 and Zadoks 10 stages of barley shoot development. Tight DNA-protein complexes of the nuclear matrix and those detected by NPC-chromatography were revealed as also involved in tissue- and development-dependent transitions, however, in sites different from TBP-DNA interactions. The spectrum of TBPs appeared to be organ and developmental-stage specific. Development of the first leaf and root system (from Zadoks 07 to Zadoks 10 stage) was shown as followed by a drastic increase in the TBP number in contrast to coleoptile, where the TBPs spectrum became poor during senescence. It was demonstrated that a nuclear protein of low molecular weight similar to the described TBPs possessed a high affinity to the DNA involved in TBP-DNA complexes.
Conclusion: Plant development is followed by redistribution of TBP along individual genes and chromosomes.
Figures
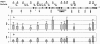



Similar articles
-
Distribution of tight DNA-protein complexes along the barley chromosome 1H, as revealed by microsatellite marker analysis.Cell Mol Biol Lett. 2005;10(1):49-59. Cell Mol Biol Lett. 2005. PMID: 15809679
-
Epigenetic chromatin modifiers in barley: IV. The study of barley polycomb group (PcG) genes during seed development and in response to external ABA.BMC Plant Biol. 2010 Apr 21;10:73. doi: 10.1186/1471-2229-10-73. BMC Plant Biol. 2010. PMID: 20409301 Free PMC article.
-
Phosphatase activity in barley proteins tightly bound to DNA and its development-dependent changes.Biochemistry (Mosc). 2012 Jun;77(6):679-88. doi: 10.1134/S0006297912060168. Biochemistry (Mosc). 2012. PMID: 22817469
-
Seed development and differentiation: a role for metabolic regulation.Plant Biol (Stuttg). 2004 Jul;6(4):375-86. doi: 10.1055/s-2004-817908. Plant Biol (Stuttg). 2004. PMID: 15248120 Review.
-
Tightly bound to DNA proteins: possible universal substrates for intranuclear processes.Gene. 2012 Jan 15;492(1):54-64. doi: 10.1016/j.gene.2011.09.019. Epub 2011 Oct 6. Gene. 2012. PMID: 22001404 Review.
Cited by
-
Tight DNA-protein complexes isolated from barley seedlings are rich in potential guanine quadruplex sequences.PeerJ. 2020 Feb 18;8:e8569. doi: 10.7717/peerj.8569. eCollection 2020. PeerJ. 2020. PMID: 32110488 Free PMC article.
References
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Research Materials
Miscellaneous

